Determining an appropriate treatment through precision medicine requires inspection of crucial novel biomarkers from tissue samples or liquid biopsies such as plasma/serum, sputum, saliva, urine, and cerebrospinal fluid for earlier disease diagnosis and prognosis. Analysis of non-coding RNA such as microRNA (miRNA) and long noncoding RNA (lncRNA), DNA methylation, and posttranslational histone modifications are epigenetic variables that have been used for cancer diagnosis, prognosis, and monitoring response.
Long noncoding RNAs (lncRNAs) are groups of transcripts composed of at least 200 non-protein-coding nucleotide sequences either from protein-coding loci or from intergenic regions. They were previously categorized as “transcriptional junk”. However, evidence showed their roles in regulatory functions where their dysregulation could spell an impending initiation and progression of human cancers. Studies showed that they actively participate in mediating stem cell functions, therefore, adding complexity to gene regulation. While their significance in normal stem cells has been proven, little attention is given to their role in cancer stem cell regulation. Tissue homeostasis relies on the spectrum of adult stem cell activity. The role of lncRNAs in stem cells mainly features maintenance; however, their role in stem cell renewal and differentiation, which is already established in microRNAs, is yet to be studied in detail.
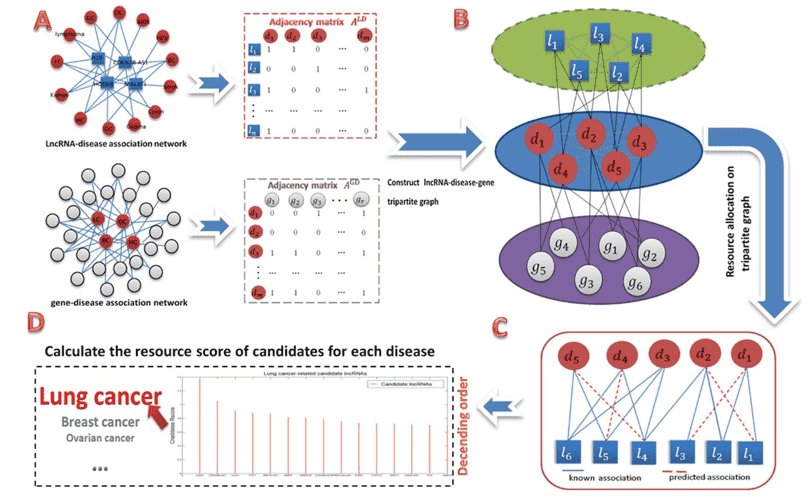
Figure 1. The screening of lncRNA candidates that are involved with lung cancer. (Ding, 2018)
LncRNAs are shown to be poorly conserved. Like rRNA and tRNA, lncRNAs operate through the formation of secondary structures. Unlike proteins, SNPs and frameshift mutations in lncRNA transcripts are less likely to negatively affect lncRNA genes since only small segments of the transcripts are needed to form 3D structures for proper functioning. Compared to protein-coding regions, compensatory mutations are relatively more common in preserving lncRNA secondary structures.
In cancer stem cells, lncRNAs are found to be currently involved in proliferation, migration, metabolism, senescence, autophagy, invasion, apoptosis, differentiation, and pluripotency. As shown in the table, they form associations creating a regulatory function of relevant proteins, mRNAs, and microRNAs.
List of lncRNA transcripts and their respective molecular associations and effects (Mauger, 2019)
|
LncRNA transcript
|
Molecular associations
|
Effect of Regulatory Function
|
|
LINC00675
|
Increases phosphorylation of vimentin
|
Suppresses gastric cancer metastasis
|
|
LncRNA HCAL
|
Blocks miR-196a/b-mediated LAPTM4B suppression
|
Enhances growth and metastasis in liver cancer
|
|
FAL1
|
Stabilizes BMI1 protein
|
Modulate the CDKN1A expression and tumor growth
|
|
LncRNA CASC11
|
Increases stability of snail mRNA
|
Promotes osteosarcoma metastasis
|
|
Lnc-DILC
|
Suppresses the IL-6/STAT3 signaling via inactivating IL-6 transcription
|
Inhibits tumorigenesis and metastasis of liver and colorectal cancer
|
|
Activates Wnt/β-catenin pathway
|
Facilitates tumorigenicity and metastasis of gallbladder cancer cells
|
There could be some instances that a lncRNA transcript could either facilitate or inhibit metastasis and tumor growth depending on certain molecular interactions. For instance, lnc-DILC or lncRNA downregulated in liver cancer stem cells acts as a tumor suppressor to inhibit the tumor growth and metastasis in liver and colorectal cancer by suppressing the IL-6/STAT3 signaling via IL-6 transcription inactivation. Conversely, lncDILC could be also upregulated in gallbladder cancer facilitating metastasis and tumor initiation of gallbladder cancer cells via Wnt/β-catenin pathway activation.
LncRNA hybridization methods and protein/ RNA immunoprecipitation are the two approaches developed to identify lncRNA-protein interactions. Specifically, these are ChIRP, CHART, RAP, CLIP, HITS-CLIP, and PAR-CLIP. Moreover, the identification of new lncRNA biomarkers is performed by microarrays and RNA-seq. RT-qPCR and digital PCR can analyze a limited panel of probed lncRNA and quantify low-abundance lncRNA.
References:
- Mauger F, Deleuze JF. Technological advances in studying epigenetics biomarkers of prognostic potential for clinical research. InPrognostic Epigenetics. 2019.
- Ding L, Wang M, Sun D, et al. TPGLDA: Novel prediction of associations between lncRNAs and diseases via lncRNA-disease-gene tripartite graph. Scientific reports. 2018, 8(1):1-1.
- Yang T, Shi Y, Yildirim E. Implications of Long Noncoding RNAs in Cancer Epigenetics. InCancer and Noncoding RNAs. 2018.
- Zhang H, Wei P, Lv W, et al. Long noncoding RNA lnc-DILC stabilizes PTEN and suppresses clear cell renal cell carcinoma progression. Cell & bioscience. 2019, 9(1):81.


 Sample Submission Guidelines
Sample Submission Guidelines