With the advent of the post-genome era, gene chip and next-generation sequencing have become two important branches of biochemistry and molecular biology technologies.
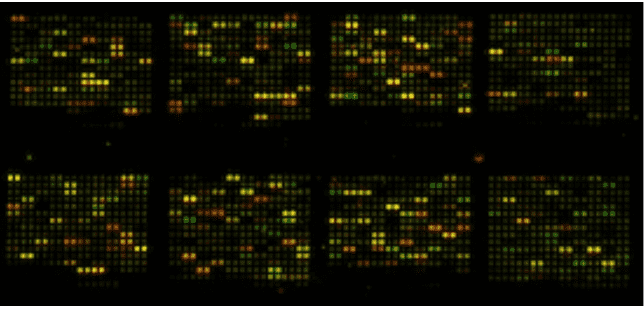
DNA microarray, also known as gene chip, is a method characterized by high throughput, high efficiency, and high automation. In a gene chip array, a large number of known probe sequences are orderly attached to a solid surface. Fluorescent molecules are labeled on the DNA to be tested, which are then hybridized with the fixed probes. According to the amount of hybridization between the tagged DNA and the probes, the fluorescence intensity varies, which is converted into data that indicates gene information. DNA microarray technology allows for the investigation and analysis of large-scale genes in a single reaction and addresses issues more straightforwardly in gene expression.
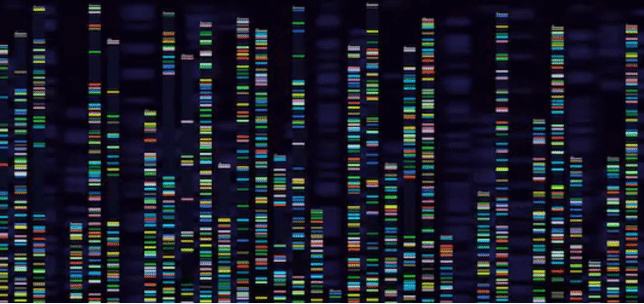
High-throughput sequencing (HTS) has been developed since the 21st century. Continued improvement has made it into an efficient, accurate, and versatile technology. The rapid development of HTS is attributed to the fact that it does not need the construction of probe library in the early stage and data analysis is conducted on mature bioinformatics platform, which alleviates most of the work and avoids the error of subcloning, and the results are more accurate. In addition, HTS can provide a technical route for polymorphism analysis of highly repetitive genomes and also plays a prominent role in the detection of DNA methylation.
Gut microbiota is a complex research field that studies the relationship between microorganisms and their host, predominantly analyzing the abundance, composition, and activity of microbial communities.
Applications in Gut Microbiome Research
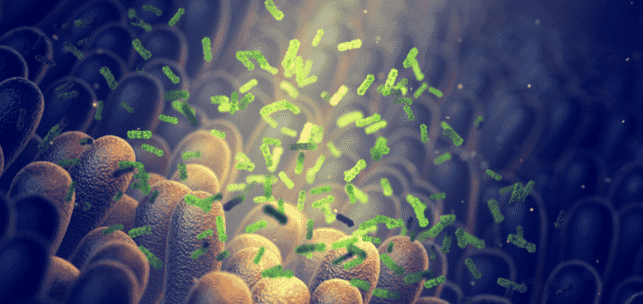
The human gastrointestinal tract is colonized by a large number of microbes including bacteria, archaea, microbial eukaryotes, and viruses, which form communities that are specific to people of different ages, diet habits, and health conditions. The gut microbiota is deemed to play pivotal roles in maintaining host health, as well as involved in many pathogenesis procedures.
DNA microarray technology is a convenient, efficient, and affordable method for routine studies of targeted intestinal microbial sequences. In a DNA microarray, a DNA library is established from already known and studied sequences, which can be applied in various research directions, including monitoring community composition and variance, functional gene analysis, physiological alterations of gut microbiota, and phylogenetic analysis. The drawback of a broad microarray is the higher number of false positives due to the vast parallel hybridization. This can be alleviated to some extent through a rigorous probe selection process and strict criteria for the positive detection of calls. Meanwhile, community-specific microarrays benefit from reduced hybridization potential, but can only be applied to the analysis of particular communities.
High-throughput sequencing technologies are getting cheaper, easier, and more powerful, hence becoming readily accessible for many researchers to study the gut microbiota. The development of high-throughput sequencing technology has greatly promoted the study of gut microbiota. The cloning step that was necessary for microarray is no longer required. And greater yields of sequencing data can be obtained from HTS. The HTS technology can analyze the composition and diversity of the microbial community, and also provide information for understanding the functions encoded by the genomes of the gut microbiota.
The above two techniques have benefits of their own. Application of either technology can address different research needs. Meanwhile, combining them can expand the interpretation of genetic information.


 Sample Submission Guidelines
Sample Submission Guidelines