Next-generation sequencing (NGS), the state-of-the-art technology that enables deep, high-throughput, and in-parallel DNA sequencing, has dramatically brought the cost down and reduced sequencing time by using massively parallel processing. NGS allows for the taxonomic assignment of microbial species in complex microbial communities, evolutionary studies, network analysis, and correlation analysis. With the efficiency and the decreasing cost, NGS has been introduced into routine laboratory practice and is particularly suitable for the analysis of microorganisms present in a given environment. NGS has revolutionized the study of genomics and molecular biology over the last two decades and brought to numerous innovative ideas and interesting applications.
Amplicon-Based Next Generation Sequencing
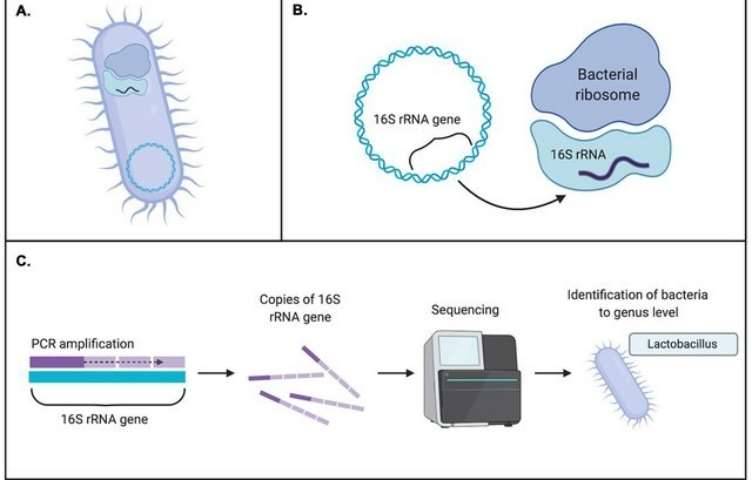
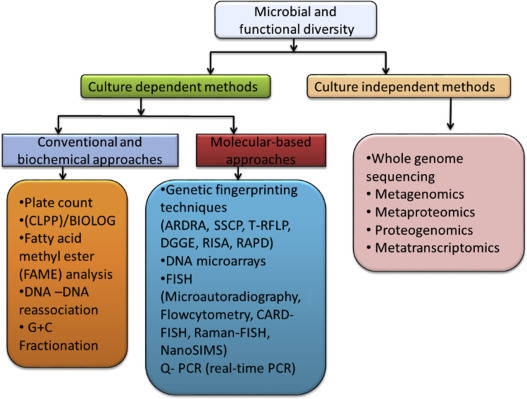
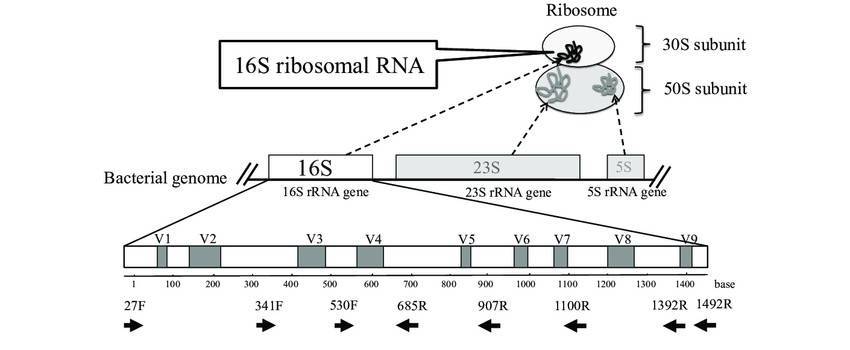
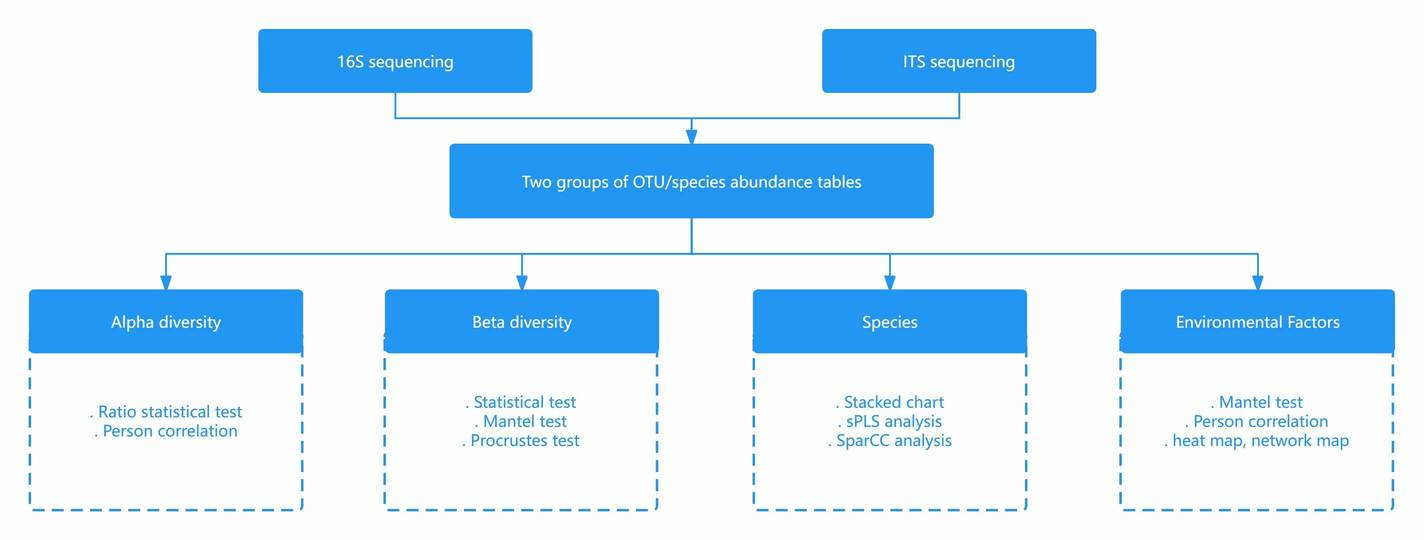
Amplicon-based NGS technologies have enabled fast, accurate, and cost-effective metataxonomic detection. The versatile amplicons of NGS provide flexible solutions to meet the needs of different research projects. Ribosomal RNA contains multiple conservative and variable regions that provide excellent genetic material for metataxonomic amplicon profiling. 16S, 18S, and ITS are well-studied and frequently targeted regions in amplicon-based NGS. Within the frame of NGS technologies, 16S rRNA sequencing is widely used in phylogenetic and taxonomic studies of bacteria and archaea. 18S rRNA sequencing can be applied to microbiome analysis of microbial eukaryotes such as fungi and protists. ITS sequencing is also a preferred method of fungal ecology studies, which is of great significance to the molecular phylogeny of fungi at the level of genus and species. Custom amplicon sequencing can further realize the pluripotency of this promising technology, by which the targeted regions are not necessarily restrained within conventional 16S/18S/ITS, providing highly specific genetic information to meet the needs of different projects.
Metagenomic Shotgun Sequencing
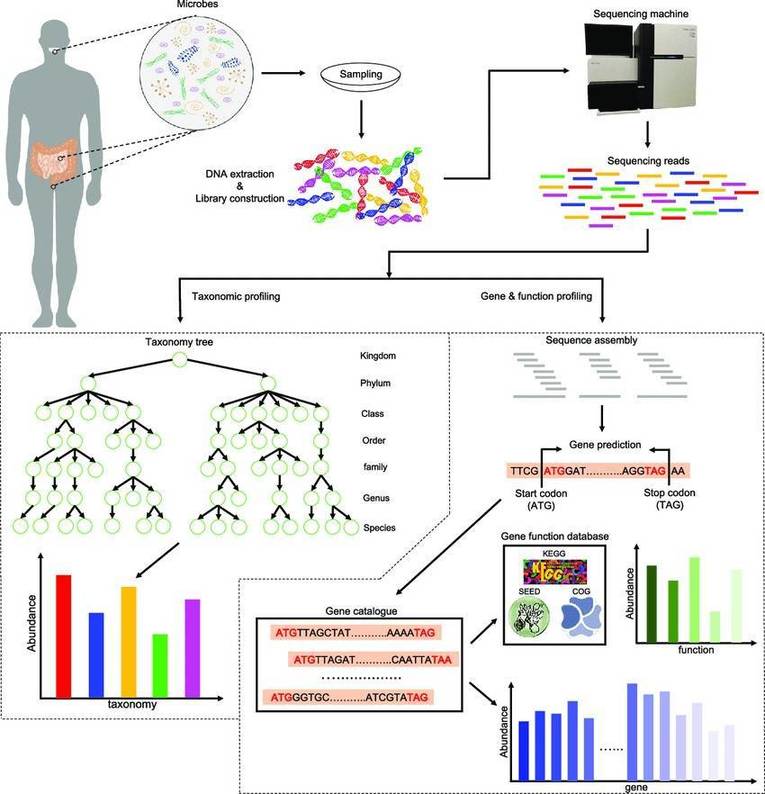
Metagenomic shotgun sequencing, which overcomes many of the limitations of amplicon sequencing, is an unbiased sequencing technology that detects pre-fragmented billions of DNA base pairs in a single run. Instead of only concerning about the targeted regions as amplicon-based sequencing, metagenomic shotgun sequencing independently sequences all DNA fragments that are randomly sheared in analogy with the pattern of a ‘shotgun’. Some of these reads will be generated from taxonomically informative regions such as 16S, 18S, and ITS1/2, while others will be generated from coding sequences to gain insight into the biological functions encoded in the genome, which simultaneously provides insight into biodiversity and function of a microbial community.
![]() metagenomics and amplicon based metataxonomic sequencing">Figure 1. The difference between shotgun metagenomics and amplicon based metataxonomic sequencing (Sekse 2017)
metagenomics and amplicon based metataxonomic sequencing">Figure 1. The difference between shotgun metagenomics and amplicon based metataxonomic sequencing (Sekse 2017)
A Comparison of Amplicon-based Next-generation Sequencing and Metagenomic Shotgun Sequencing
| Amplicon-Based Next Generation Sequencing | Metagenomic Shotgun Sequencing | |
|---|---|---|
| Principle | Amplicon-based NGS uses oligonucleotide probes designed to target and capture regions of interest – generally hypervariable regions of conserved genes or intergenic regions, followed by next-generation sequencing to provide genetic information of these amplified sequences. | Metagenomic shotgun sequencing involves randomly shearing the DNA of the microbial genome into small fragments, then adding a universal primer at both ends of the fragments for PCR amplification and sequencing. And then splice the sequenced of the small fragments into a longer sequence through assembly. |
| Research Objectives | Amplicon-based NGS assays are designed mainly for the purpose of studying the phylogenetic relationship of species, the species composition, and the biodiversity of a microbial community. | Apart from the taxonomic analysis that amplicon-based NGS can provide, metagenomic shotgun sequencing can also conduct in-depth research on genes and functions of a microbial community, such as pathway analysis, KEGG, GO, etc. |
| Taxonomic Resolution | Microorganisms can be reliably identified at the genus levels, and some at species levels. | Metagenomic shotgun sequencing can identify microbes at the species levels and even provides discrimination power for subspecies and strains. |
| Advantages |
|
|
| Disadvantages |
|
|
| Recommended Applications | Evaluating the differences in a large number of microbiota samples across different environments | Deeply investigating a smaller number of samples |
Due to the respective advantages of amplicon-based NGS and metagenomic shotgun sequencing, as well as the different databases they referred to, the two approaches have their own specialties and can be combined to study the composition, diversity and functions of microbial communities, delivering an accurate and comprehensive result.
Reference
- Sekse C; et al. High throughput sequencing for detection of foodborne pathogens. Frontiers in microbiology. 2017, 8:2029.