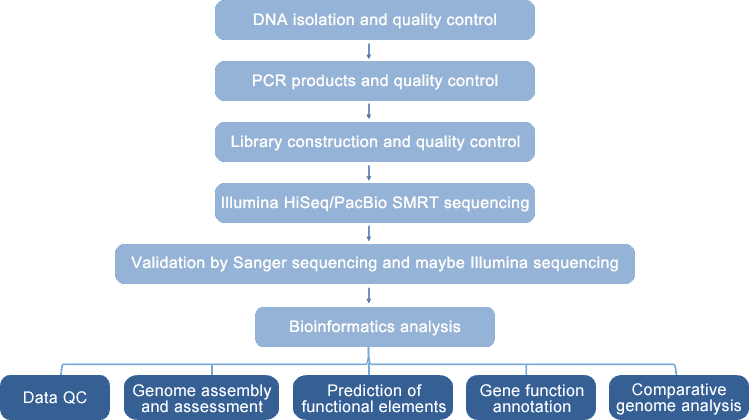
The automated and excellent bacterial whole genome resequencing platform at CD Genomics enables high-effective and affordable microbial whole genome resequencing. We perform PacBio SMRT (100-150X) or Illumina HiSeq PE150 (300-500 bp insert DNA library, 50-100X) sequencing with your samples. PacBio SMRT sequencing generates long reads and Illumina sequencing is characterized by high throughput and high accuracy rate. After raw data QC and filtering, qualified reads are aligned to the reference genome for detection of variants, including single nucleotide polymorphisms (SNPs), insertions/deletions (InDels), structure variations (SVs), and copy number variations (CNVs). These molecular markers can further be functionally annotated and used for studies of evolution, population characterization, and selective pressure.

Unlike PCR-based or capillary sequencing approaches, next-generation sequencing (NGS)-based microbial whole genome resequencing allows researchers to sequence hundreds of organisms with the power of multiplexing. At the same time, it simplifies the workflow and saves time. NGS can identify low-frequency variants and genome rearrangements in an efficient and economical way.
Our bacterial whole genome resequencing platform enables researchers to conduct research on disease outbreaks and treatments, as well as drug development and microbial control in many fields. If your microbial samples have no reference genomes, we also have bacterial whole genome de novo sequencing platform that generates accurate reference genomes, which are useful for microbial identification and comparative genomic studies.