Microbial Therapeutics: New Opportunities for Medicine
Microorganisms were thought to cause serious disease. However, some types of the microorganisms can synthesize biologically active molecules and be used as therapeutics. More pharmaceutical companies have realized the values and directed their clinical development efforts towards microbial therapeutics. The potential of microbial therapeutics is evident in the probiotics and live bacterial prophylactic vaccines, as well as more the recent cases using fecal microbiota transplant to improve human health or cure certain diseases. Besides, microorganisms can be engineered as live therapeutics to perform specific functions in the human body. Benefits could include preventing infections, eliminating tumors, and treating metabolic disorders. Engineered living systems are considered the next generation of therapeutics. Producing therapeutics derived from the microbes has several advantages over traditional systems: decreased production costs, side effects, and dosage requirements.
Our Solution for Microbial Therapeutics
The microbiome has progressively become an attractive target for potential therapeutics. However, one of the biggest difficulties is to determine cause-effect relationships and to design microbial therapeutics for these conditions. We are dedicated to offering a full list of microbial genomics services to accelerate the development of microbial therapeutics, including microbial whole genome sequencing, 16S/18S/ITS sequencing, metagenomics, metatranscriptomics, microbial epigenomics, prokaryotic RNA sequencing, as well as microbial ID and AMR profiling.
Main Research Directions
- Reveal the relationship between microbes and human disease.
- Development of engineered microbes to cure certain human diseases.
- Microbiome-based therapies to alter the microbial composition of human body, like the fecal microbiota transplant.
What Can We Do?
- 1. Microbial diversity analysis of the human microbiome on the skin, or in the gut.
- 2. Metabolomic pathway characterization of the microbial ecosystem.
- 3. Comprehensive bioinformatics analysis to define functional signatures for disease states.
- 4. Testing and validation of engineered microorganisms.
- 5. Microbiological Quality Control (QC) in the Pharmaceutical Industry.
Note: Our service is for research use only, not for disease diagnosis or treatment.
Detectable Objects
Bacteria, archeae, fungi, protozoan, viruses.
Detection Methods
Next-generation sequencing (NGS), long-read sequencing, quantitative PCR (qPCR), and other molecular biology techniques.
Detection Platforms
Illumina HiSeq/MiSeq, Nanopore systems, PacBio RSII, PacBio Sequel, Applied Biosystems real-time PCR platforms, et al.
Sample Requirement
-

- DNA sample: 1.8 < OD260/280 < 2.0 (without degradation or contamination)
- RNA sample: RIN ≥ 7.0, 28s/18s ≥ 1.5 (without degradation or contamination)
Workflow
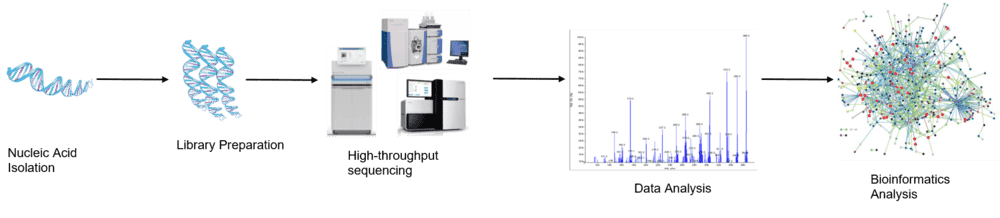 Figure 1. The high-throughput sequencing analysis process.
Figure 1. The high-throughput sequencing analysis process.
Bioinformatics Analysis
Our bioinformatics analysis pipeline is flexible to your needs.
| Taxonomic Assignment | Species annotation |
| Rarefaction Curve | |
| Shann-Wiener Curve | |
| Rank Abundance Curve | |
| Statistical Analysis | Heatmap |
| VENN | |
| Principal Components Analysis (PCA) | |
| Comparative Analysis | RDA/CCA |
| PCA/PCoA | |
| Network Analysis | |
| Functional analysis | BLASTX, KEGG, eggNOG, CAZy, CARD, ARDB etc. |
| PICRUSt | |
| TAX4Fun | |
| Phylogenetic Analysis | (Un)Weighted Unifrac |
| Phylogenetic Trees |
References
- Mimee M, Citorik R J, Lu T K. Microbiome therapeutics—advances and challenges. Advanced drug delivery reviews, 2016, 105: 44-54.
- Pedrolli, et al. Engineering Microbial Living Therapeutics: The Synthetic Biology Toolbox. Trends in Biotechnology, 2018, doi:10.1016/j.tibtech.2018.09.005.