| Genome Analysis |
Genome assembly |
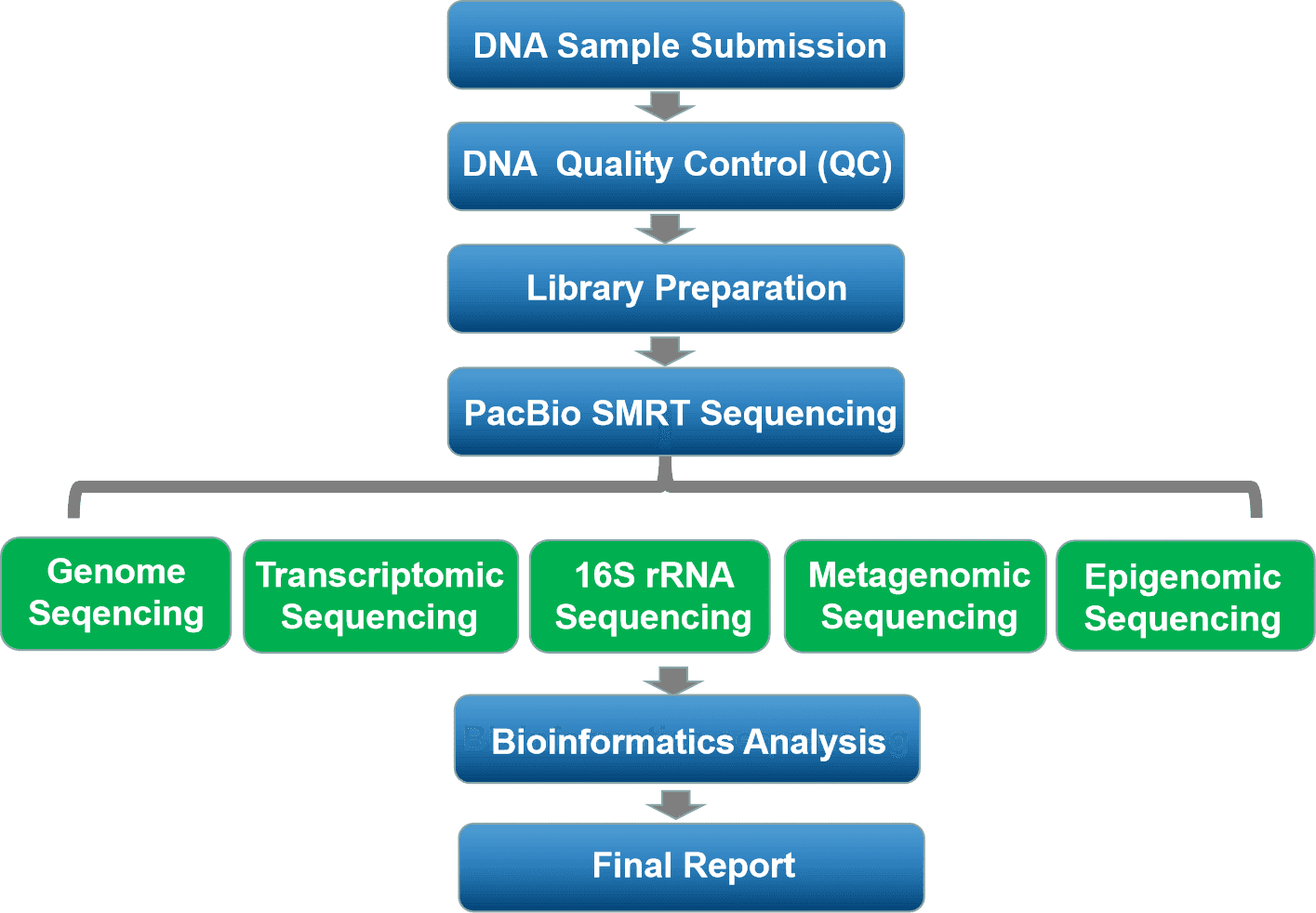
High-quality de novo assemblies of genomes using Hierarchical Genome Assembly Process (HGAP) |
| Functional annotation |
COG, KEGG, GO, etc. |
| Genome annotation |
Prediction of Non-coding RNAs, detection of CRISPRs, etc. |
| Evolutionary analysis |
Construction of phylogenetic trees |
| Variant calling |
Detection and annotation of copy number variations (CNVs), structural variations (SVs), single nucleotide polymorphisms (SNPs), and insertion-deletions (InDels) |
| Transcriptomics Analysis |
Isoform analysis |
Reference genome mapping, full-length isoform correction and classification, reduced redundancy |
| Functional annotation |
GO,KEGG, KOG, COG, Swissport, etc. |
| Gene expression analysis |
Analysis of gene abundance, identification of differentially expressed genes (DEGs) |
| Structure identification |
Prediction of alternative splicing events, LncRNAs, miRNAs, and CDS, the discovery of novel transcripts, identification of fusion genes, etc. |
| SNPs/SSRs analysis |
Detection and annotation of SNPs and simple sequence repeats (SSRs) |
| 16S rRNA Sequencing Analysis |
OUT clustering |
Clustering and filtering of operational taxonomic units (OTUs) |
| Taxonomic assignment |
Venn diagram, rank abundance curve, PCA, etc. |
| Diversity analysis |
Alpha diversity, beta diversity, meta analysis |
| Functional prediction |
PICRUSt, FAPROTAX, Tax4fun, FunGuild |
| Metagenomics |
Assembly |
Reference-based assembly, de novo assembly, and quality assessment of metagenomics assemblies. |
| Taxonomic assignment |
Taxonomic assignment, species abundance analysis |
| Epigenomics functional annotation |
Metagenome gene prediction, protein function prediction, and pathway annotation |
| Custom Analysis |
More data mining upon your request |