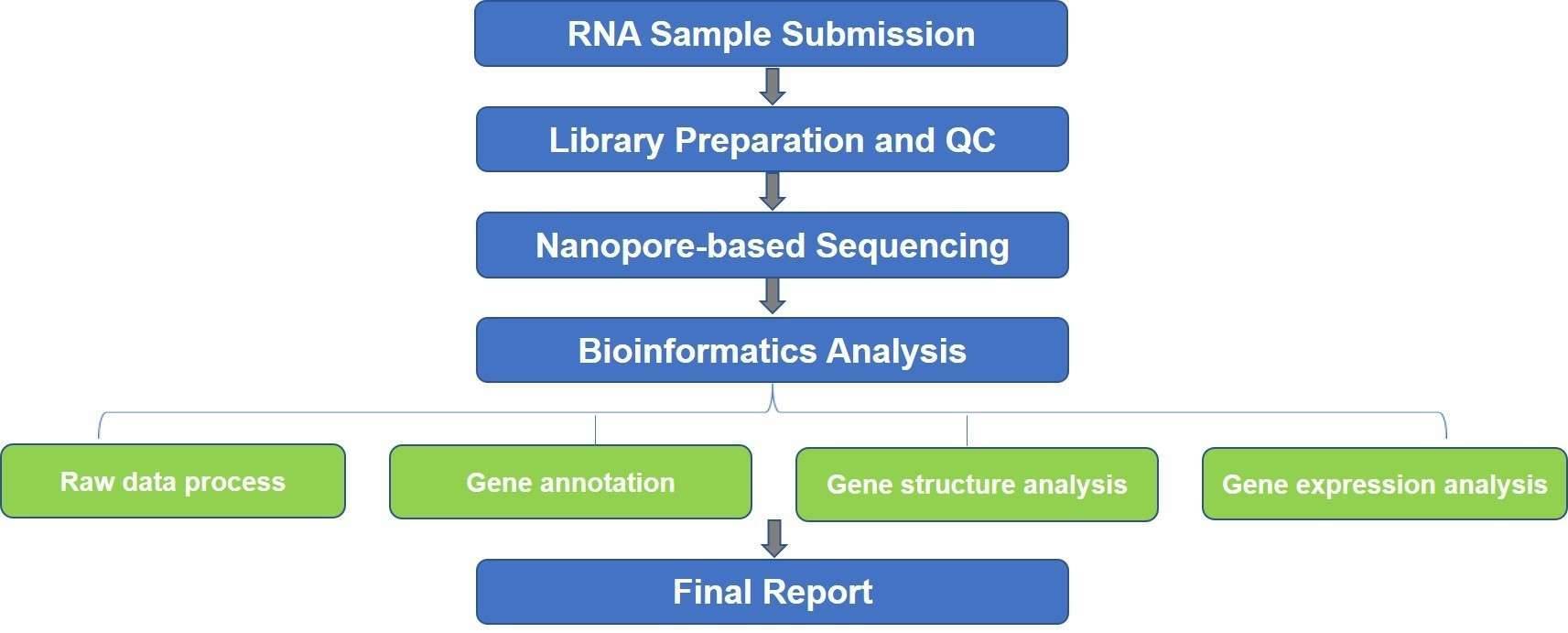
The study of microbial transcriptome and metatranscriptome is important for detecting RNA isoforms, quantifying gene expression levels, predicting resistance to specific antibiotics, understanding host-pathogen immune interactions, and tracking disease progression. Our nanopore-based microbial transcriptomics enables a relatively simple workflow and provides full-length, highly accurate transcriptome data of bacteria, archeae, fungi, and viruses, so as to reveal the complete set of mRNAs, circular RNA (circRNA), long non-coding RNA (lncRNA), microRNA (miRNA), and other non-coding RNAs in a sample under specific conditions. Nanopore sequencing enables direct RNA sequencing and simultaneous detection of epigenetic RNA modifications.
Microbial transcriptomics analysis can provide a snapshot of actively expressed genes and transcripts under specific conditions. It plays an importance role in gene annotation, isoform identification, identification of fusion transcripts, and lncRNA identification. It can also be used to elucidate the molecular mechanisms and biological pathways that regulate microbial activities, and to study host-microbe interactions like the effects of host environments on microbial gene expression and metabolic pathways.