Pathogen Targeted Sequencing, also known as tNGS, is a targeted capture-based next-generation sequencing (NGS) technology. It operates independently of traditional microbial cultivation methods, directly enriching nucleic acids from clinical samples. Subsequently, high-throughput sequencing is performed, followed by alignment and analysis against a database. Based on the aligned sequence information, the types of pathogenic microorganisms present in the sample can be rapidly and objectively detected. This includes viruses, bacteria, fungi, parasites, and more. tNGS offers advantages such as a well-defined pathogen spectrum and cost-effective sequencing.
Pathogenic tNGS Service
CD Genomics Launches Comprehensive Pathogenic Sequencing Solution, expanding the scope of detection, optimizing cost-effectiveness, enhancing efficiency, and maximizing productivity. This innovative solution aims to continually elevate sequencing capabilities, contributing to the development of a robust sequencing ecosystem.
Applicable Sample Types
Peripheral blood, bone marrow, sputum, bronchoalveolar lavage fluid, cerebrospinal fluid, other puncture fluids, urine, ocular, aural, oral, nasal, and throat secretions, genital secretions, wound secretions, pus, tissues, and various clinical infection specimens.
Pathogen Coverage
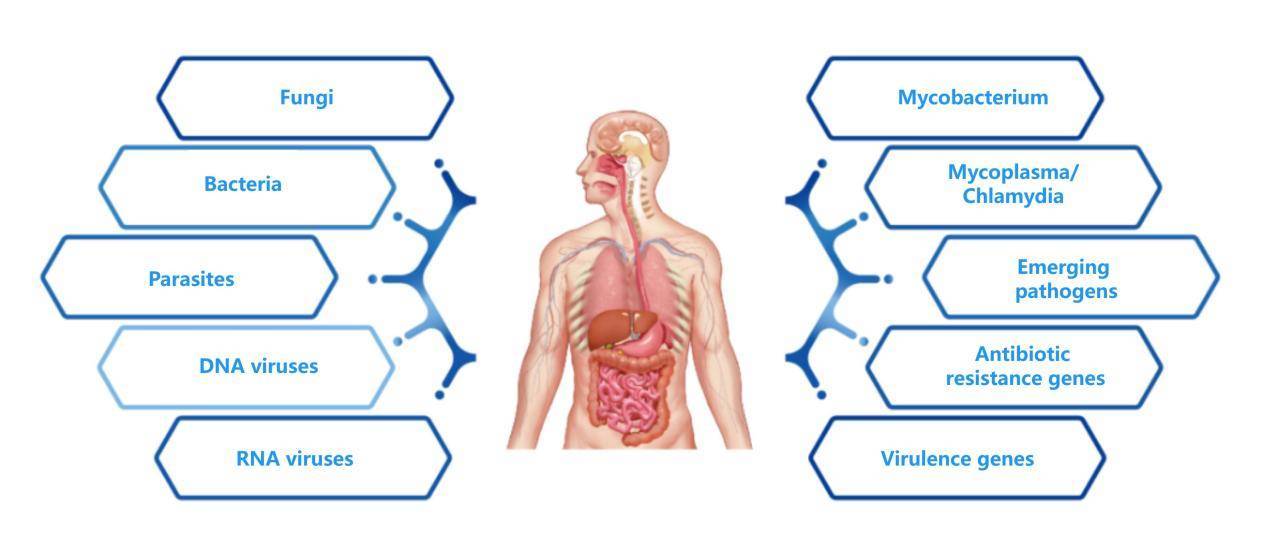
Encompassing 200+ pathogenic microorganisms, 80+ drug-resistant genes, and 40+ virulence genes, it addresses common pathogens in various clinical infection scenarios, including bloodstream infections, central nervous system infections, respiratory tract infections, and joint infections.
Product Advantages
Covers over 95% of clinically infectious pathogens with high incidence.
Achieves sensitivity and specificity in detection both exceeding 90%.
Unaffected by contaminating microbes and host sequences.
Can tNGS Replace mNGS?
tNGS and mNGS each exhibit distinct advantages and limitations, making them applicable in different scenarios. The strength of tNGS lies in its ability to detect low-abundance pathogens with high sensitivity while reducing the sequencing data volume and analysis workload. This enhances detection efficiency and cost-effectiveness. However, tNGS is constrained by its inability to identify novel or rare pathogens and the limitation of only detecting pre-designed pathogens. Furthermore, it cannot conduct whole-genome analysis of pathogens.
On the other hand, mNGS excels in unbiased detection of all pathogens within a sample, enabling the discovery of novel or rare pathogens. It also provides comprehensive genomic information on pathogens, facilitating traceability, genotyping, and drug resistance assessment. Nevertheless, mNGS demands substantial sequencing data and analysis efforts, resulting in higher detection costs and time consumption. It exhibits lower sensitivity in detecting low-abundance pathogens.
Therefore, tNGS and mNGS are not mutually exclusive; instead, they complement and collaborate with each other. In the field of clinical microbiology, tNGS is more suitable for rapid screening and diagnosis of common or known pathogens, while mNGS is better suited for exploring and identifying unknown or rare pathogens. Integrating both technologies enables a more comprehensive, accurate, and efficient detection of pathogens.
In addition, CD Genomics leverages NGS technology to provide metagenomic sequencing (mNGS), and comprehensive pathogen whole genomics sequencing services to global pathogen research projects.

