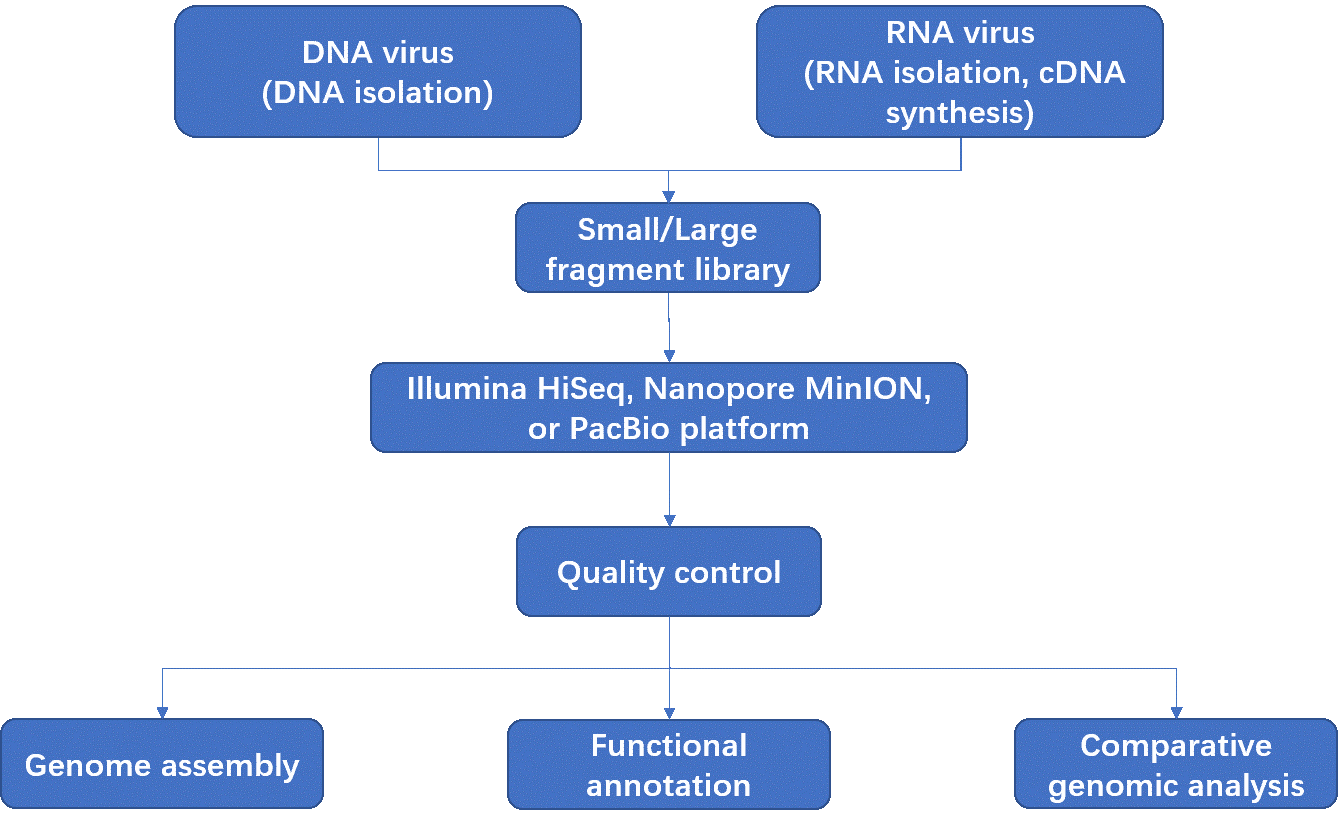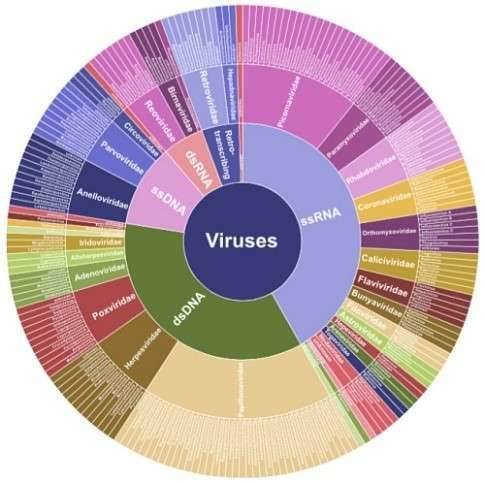
Whole-genome sequencing of pathogens is becoming increasingly important for both basic research and applications. Viral genome resequencing refers to the sequencing of all genetic information of viruses in a time-saving and cost-effective way, identifying sequence variants and providing insight into viral phylogeny and evolution, and pathogenesis. With the availability of reference sequences, reads trimming & filtering and genome assembly become more efficient and accurate. We have established standardized viral genome resequencing platform based on NGS, PacBio and Oxford nanopore instruments. PacBio SMRT sequencing and Oxford nanopore sequencing can generate much longer reads, and detect epigenetic modifications in a single sequencing run.
Viral genome resequencing is important to increase the power of molecular epidemiology, thus accelerating the development of novel treatments and vaccines. Although the progress of viral genome resequencing for applications has been slow, its importance has been gradually accepted, particularly for tracking disease outbreaks, identifying routes of viral transmission, and the detection of all known and novel drug-resistant variants.