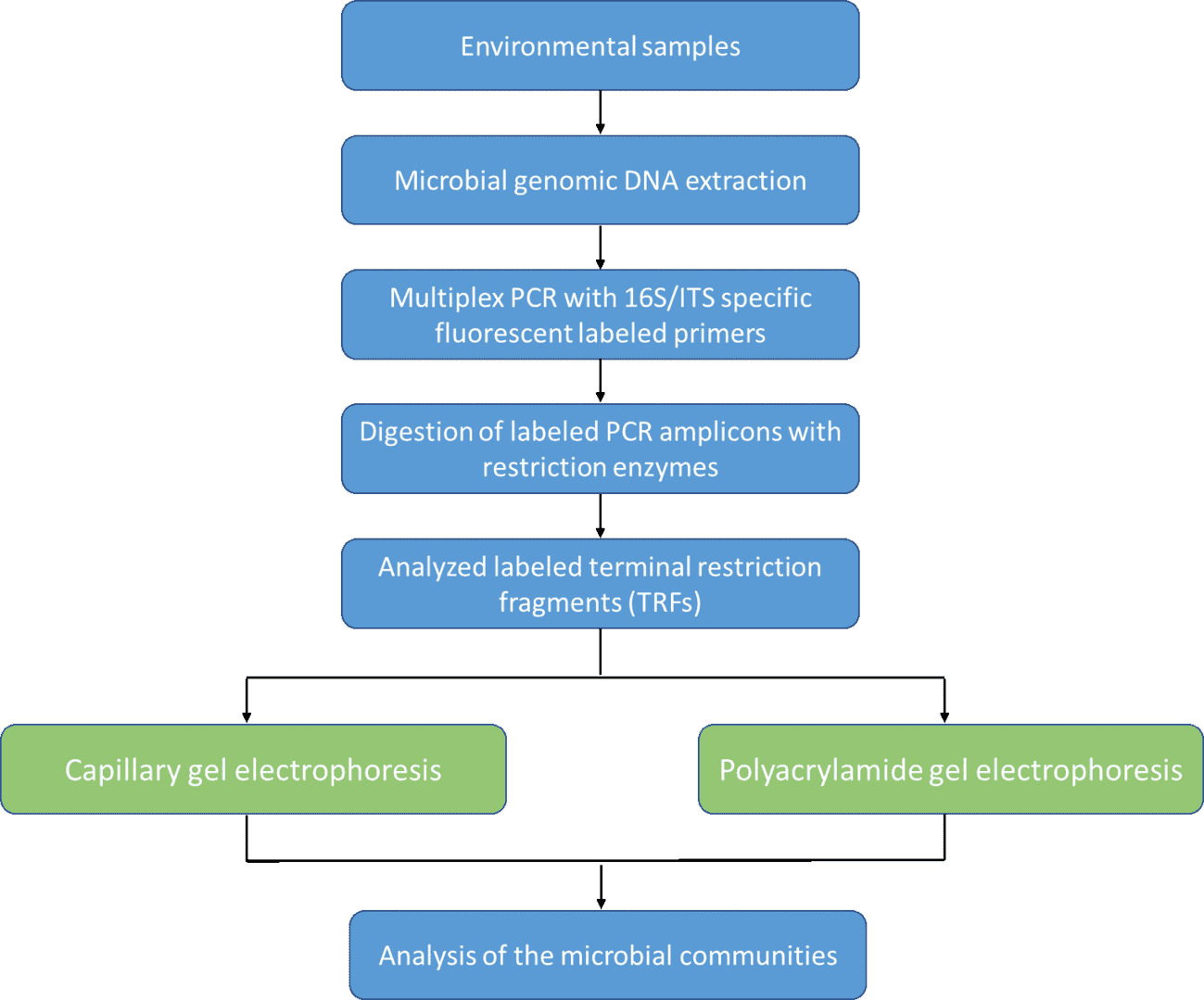
The determination of static or dynamic balances among microorganisms is of fast-growing interest. The generation of species-specific and fluorescently labeled 16S ribosomal DNA (rDNA) fragments by the terminal restriction fragment length polymorphism (T-RFLP) technique is a suitable tool that meets the criteria. For the separation of these fragments, technologies include capillary gel electrophoresis and polyacrylamide gel electrophoresis.
CD Genomics provides T-RFLP technology to fragment microbial genes, the digested amplicons are separated by polyacrylamide gel electrophoresis or by running it in capillary gel electrophoresis. Our microbial gene fragment analysis is multiplexing, allowing the assessment of more than 20 loci in a single reaction, and fragments that vary from only one base pair only are precisely sized. The automated workflows for the analysis of loads of specimens of DNA fragments within a short period.