dPCR stands for digital polymerase chain reaction, which is a method for quantifying the amount of DNA in a sample and it is a highly sensitive and quantitative PCR method.
Microbial dPCR analysis refers to the application of digital PCR to quantify the amount of microbial DNA in a sample. It works by partitioning the reaction mixture into thousands of tiny droplets, each of which acts as an independent PCR reaction. The number of positive droplets is then counted, and the amount of DNA in the original sample can be estimated from this. This method provides a highly sensitive and specific way of quantifying DNA, and is particularly useful for applications in microbial analysis, where small amounts of DNA are often present.

Tell Us About Your Project
We are dedicated to providing outstanding customer service and being reachable at all times.
 Request a Quote
Request a Quote
Applications of dPCR in Microbiological Analysis
Pathogen quantification: dPCR can be used to accurately quantify the number of bacteria, viruses, and fungi in a sample.
Antimicrobial resistance testing: dPCR can be used to detect and quantify resistance genes in bacterial pathogens.
Microbial community analysis: dPCR can be used to accurately quantify the diversity and relative abundance of microorganisms in complex microbial communities.
Gene expression analysis: dPCR can be used to quantify gene expression in microorganisms, which is important in understanding the regulation of gene expression in response to environmental conditions and other stimuli.
Overall, dPCR provides a high level of accuracy and precision for quantifying target DNA sequences in microbiological analysis, making it a valuable tool for research and clinical applications.
Workflow of Microbial dPCR Analysis
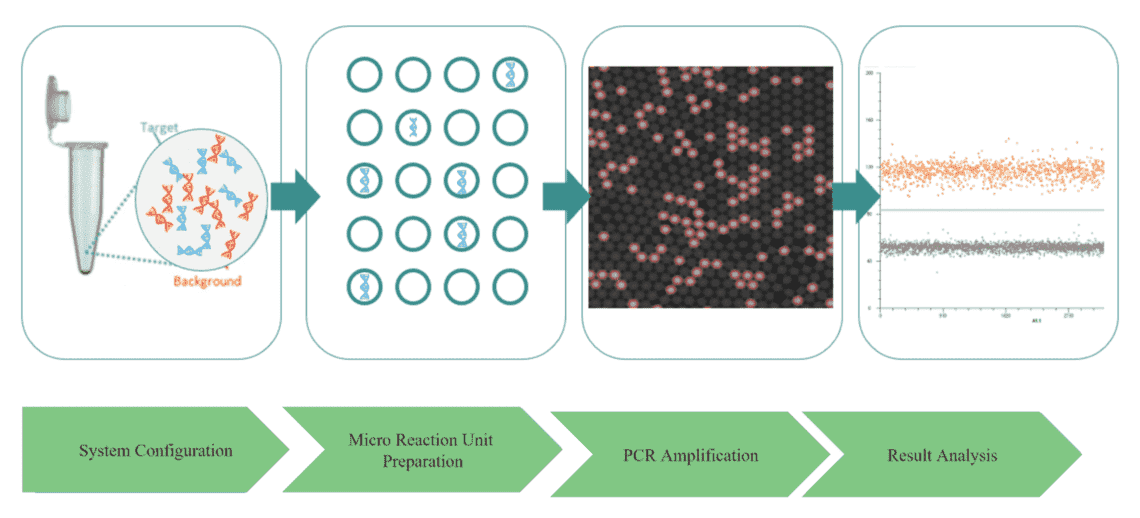
Advantages of Digital PCR
There are several advantages of Digital PCR (dPCR) compared to traditional PCR:
dPCR provides a powerful tool for a wide range of applications, including pathogen quantification, gene expression analysis, and microRNA profiling.
Increased sensitivity: dPCR has a higher sensitivity compared to traditional PCR, as it is able to detect even single DNA molecules.
Improved quantification accuracy: dPCR provides more accurate quantification of target DNA by partitioning the reaction mixture into thousands of tiny droplets, each of which acts as an independent PCR reaction. This reduces the impact of PCR inhibitors, enabling accurate quantification even in complex samples.
Increased precision: dPCR has a high degree of precision, allowing for more accurate measurement of copy number variation (CNV) and gene expression analysis.
Improved reproducibility: dPCR reduces the variability that can occur between reactions, making it a more reproducible method compared to traditional PCR.
Reduced bias: dPCR minimizes well-to-well variation that can occur in traditional PCR, leading to reduced bias in results.
Handling of low-input samples: dPCR is well-suited for the analysis of low-input samples, such as formalin-fixed paraffin-embedded (FFPE) tissue samples, that are often difficult to work with using traditional PCR.
Samples Types
Absolute amount of microorganisms such as bacteria and fungi in soil, water, feces, sediment and other samples
The absolute amount of antibiotic resistance genes in samples such as soil and water
The absolute amount of functional genes such as carbon cycle, nitrogen cycle, and sulfur cycle in samples such as soil and sediment



