The Importance of Root Microbiome Analysis
Rhizosphere microorganisms refer to the microorganisms that are closely attached to the rhizosphere soil. They are mainly bacteria, of which gram-negative bacteria including Pseudomonas, Flavobacterium, Pseudomonas, and Agrobacterium are the most predominant microorganisms. Rhizosphere microorganisms are essential for plant growth and organic matter turnover. The plant root exudates are their nutrient and energy source. They mainly affect the permeability and metabolism of root cells. CD Genomics can determine the the types and relative abundance of root microbiomes, explore the microbial diversity of the roots, and study the relationship between the root microbiome and the environment for our clients.
Accelerate Research and Practice in the Root Microbiome
16S/18S/ITS sequencing and metagenomic sequencing are ideal technologies for studying root-associated microbial diversity. According to your specific needs, we employ various sequencing platforms, including Illumina Miseq/Hiseq, Roche 454, Ion Torrent PGM, PacBio RS II, and Nanopore MinION instruments to obtain huge and accurate sequence data. Comprehensive bioinformatics approaches such as RDA, CCA, NMDS analysis are used to investigate the microbial community structure, the relationship between environmental factors (pH, C/N, humidity, total nitrogen, total phosphorus, TOC, water content, etc.) and the root microbiome, and the relationship between root microorganisms and plant growth.
Main Research Directions
- Application of single or combined beneficial rhizosphere microorganisms, such as bio-fertilizer or bio-pesticide.
- The rhizosphere microbiome and plant health.
- The rhizosphere microbiome and sustainable agricultural development.
What Can We Do?
- 1. Microbial identification and diversity analysis using 16S/18S/ITS sequencing or metagenomic sequencing.
- 2. Microbial abundance analysis using 16S/18S/ITS sequencing or real-time qPCR.
- 3. Explore soil-plant-microbe interactions and predict ecological function of microbes using sequencing technologies coupled with bioinformatics analysis.
Note: Our service is for research use only, and not for therapeutic or diagnostic use.
Detectable Objects
Bacteria, fungi, archaebacteria.
Detection Methods
Next-generation sequencing (NGS), long-read sequencing, PCR denaturing gradient gel analysis (PCR-DGGE), real-time qPCR
Technical Platforms
Illumina HiSeq/MiSeq, Ion Torrent systems, PacBio SMRT systems, Nanopore MinION systems, PCR-DGGE, Real-time qPCR, clone library, etc.
Sample Requirement
-

- DNA sample: ≥ 500 ng, OD260/280 = 1.8-2.0, concentration ≥ 10 ng/μl.
- Ensure that the DNA is not degraded. Avoid repeated freezing and thawing during sample storage and shipment.
- Please use enough dry ice or ice packs during shipment.
- Nucleic acid samples should be stored at -20 °C.
Workflow
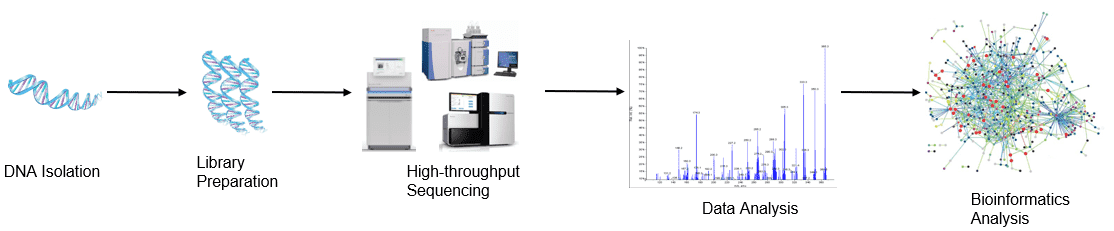 Figure 1. Our workflow for high-throughput sequencing analysis.
Figure 1. Our workflow for high-throughput sequencing analysis.
 Figure 2. Workflow for PCR-DGGE analysis process.
Figure 2. Workflow for PCR-DGGE analysis process.
Bioinformatics Analysis
| OTU Clustering | Distribution-Based OTU-Calling |
| Rarefaction Curve | |
| Shannon-Wiener Curve | |
| Rank Abundance Curve | |
| Diversity Index | |
| OTU-Based Analysis | Heatmap |
| VENN | |
| Principal Components Analysis (PCA) | |
| Hierarchical Clustering | |
| Comparative Analysis | RDA/CCA |
| PCA/PCoA | |
| Network Analysis | |
| Functional analysis | BLASTX, KEGG, eggNOG, CAZy, CARD, ARDB etc. |
| PICRUSt | |
| TAX4Fun | |
| Phylogenetic Analysis | (Un)Weighted Unifrac |
| Phylogenetic Trees |
Reference
- Ana C. C. Pires, et al. Denaturing Gradient Gel Electrophoresis and Barcoded Pyrosequencing Reveal Unprecedented Archaeal Diversity in Mangrove Sediment and Rhizosphere Samples. Applied and Environmental Microbiology, 2012, 78(16): 5520-5528.2.

