Significance of Metagenomic Resistance Gene Sequencing
The threat to human public health from the spread of ARGs has attracted the attention of scientists due to the growing problems caused by the widespread use and misuse of antibiotics. In the past decades, ARGs of different types of antibiotics, especially β-lactams, tetracyclines, aminoglycosides and sulfonamides, which are widely used in human medicine and animal farming, have been detected in humans, soil, water environment (including urban domestic and hospital wastewater, livestock farming wastewater, aquaculture environment, WWTP discharge water, rivers and sludge, etc.) and even air as well as food. The co-existence of ARGs in different environmental samples indicates that human activities lead to high abundance of ARGs in the environment and that ARGs have been widely spread in different environments. Studies have been conducted on the diversity and abundance of ARGs in different environments, transmission routes (e.g., HGT), relationship with drug resistance mechanisms, and specific microorganisms carrying specific ARGs. In the future, we will aim to explore methods to inhibit the occurrence and transmission of ARGs in different environments and prevent the transmission of ARGs to human-associated pathogens, thus reducing the threat of ARGs to human health In the future, we will explore ways to inhibit the occurrence and transmission of ARGs in different environments, prevent the transmission of ARGs to human-related pathogens, and thus reduce the threat of ARGs to human health.
In the era of rapid development of scientific information technology, the development and application of metagenomics technology has made great contributions to the research and development of human microorganisms. The development of metagenomics will move towards higher throughput, more accurate and lower cost. However, for the huge amount of data obtained from current research, how to select and construct a better metagenomics database for the screening and analysis of ARGs, as well as to interpret the content of results more clearly and simply, are still technical challenges that everyone is concerned about and need to overcome.
The culturable bacterial species currently studied by humans are still the "tip of the iceberg", and non-culture-based metagenomics is useful for dissecting the diversity and abundance of ARGs in different environments, the transmission of ARGs through MGEs, as well as the relationship between ARGs and antibiotic resistance mechanisms, and identifying the specific molecular characteristics of antibiotic resistance, which are currently the focus of many studies.
The use of metagenomics has guiding significance for the use of antibiotics in human medicine, animal breeding and disease treatment, antibiotic residues in WWTP, and the treatment of ARGs. At present, the world is facing a serious problem of antibiotic resistance, which makes the development and utilization of metagenomics particularly important. Metagenomics can explore and discover a wider range of ARGs in nature, capture a more complete ARGs map, and lay the foundation for studying the function of ARGs, predicting possible antibiotic resistance in the future, studying new antibiotics, and finding alternatives to antibiotics ; and the combination of metagenomics and other omics technologies (such as metatranscriptomics) can explore and learn more knowledge about ARGs, and provide technical support for assessing the risk of ARGs and solving the problem of antibiotic resistance in the future.



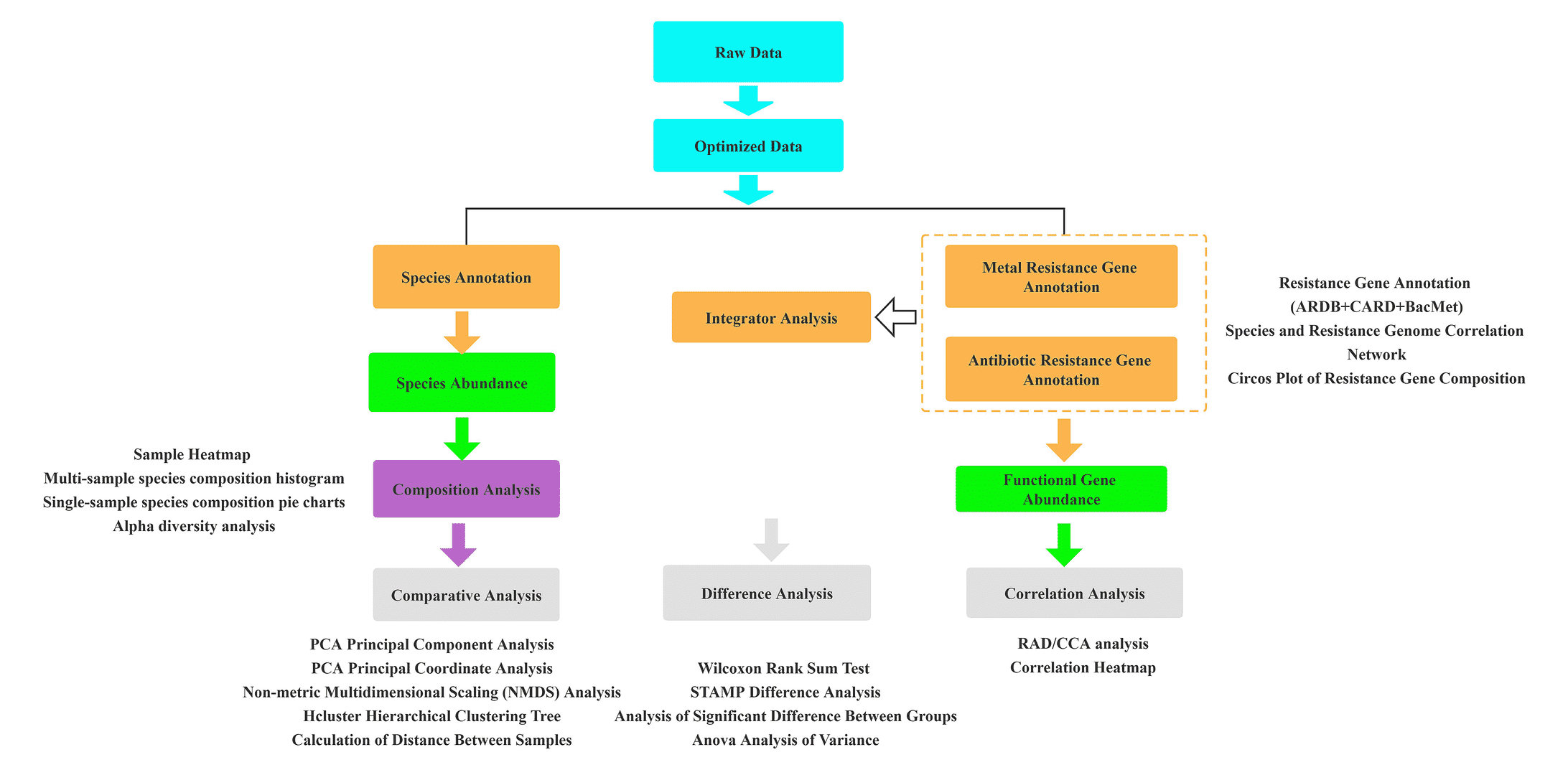 Metagenomic analysis of antibiotic resistance in microbial communities
Metagenomic analysis of antibiotic resistance in microbial communities