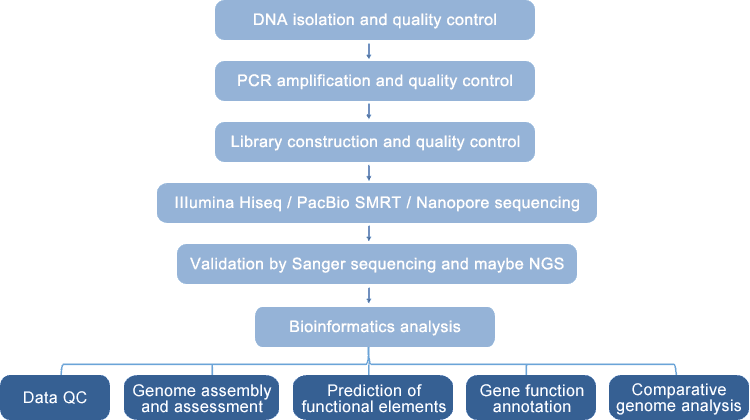
The microbial whole genome sequencing platform at CD Genomics is an automated and intelligent platform utilizing Illumina HiSeq sequencing, PacBio SMRT sequencing, and/or Nanopore sequencing. It resolves whole genome assemblies, especially for repeat-dense and GC-rich genomes, as well as plasmids. This platform can be used to sequence the complete genome of bacteria, fungi, viral/phage, archaea, algae, and protozoa by microbial de novo sequencing or resequencing. Based on the resulting genomic information, it is viable to predict the functional elements of the genome, annotate gene function, and conduct comparative genome studies, identify single nucleotide polymorphisms (SNPs) and insertions-deletions (InDels), and discover other genetic alterations among microbial strains.

Microbial whole genome sequencing further characterizes microbial methylomes. These medications may serve as a defense mechanism in microbes or regulate certain biological processes in bacteria, such as DNA replication, gene expression, DNA repair, and cell cycle. With the capability of characterizing closed microbial reference genomes and methylomes with confidence and ease, this platform has significant advantages in clarifying the role of structural variants in the evolution of virulence, and addressing issues of drug resistance and transmission paths, as well as disease outbreaks.
For different research purposes, CD Genomics has different sequencing strategies. For draft genome and microbial resequencing, we recommend Illumina HiSeq PE150 sequencing. For complete genomes, we recommend PacBio SMRT sequencing. And we ensure the 50-fold (at least) coverage of PacBio SMRT sequencing data and 100-fold (at least) coverage of Illumina Hiseq PE150 sequencing data.