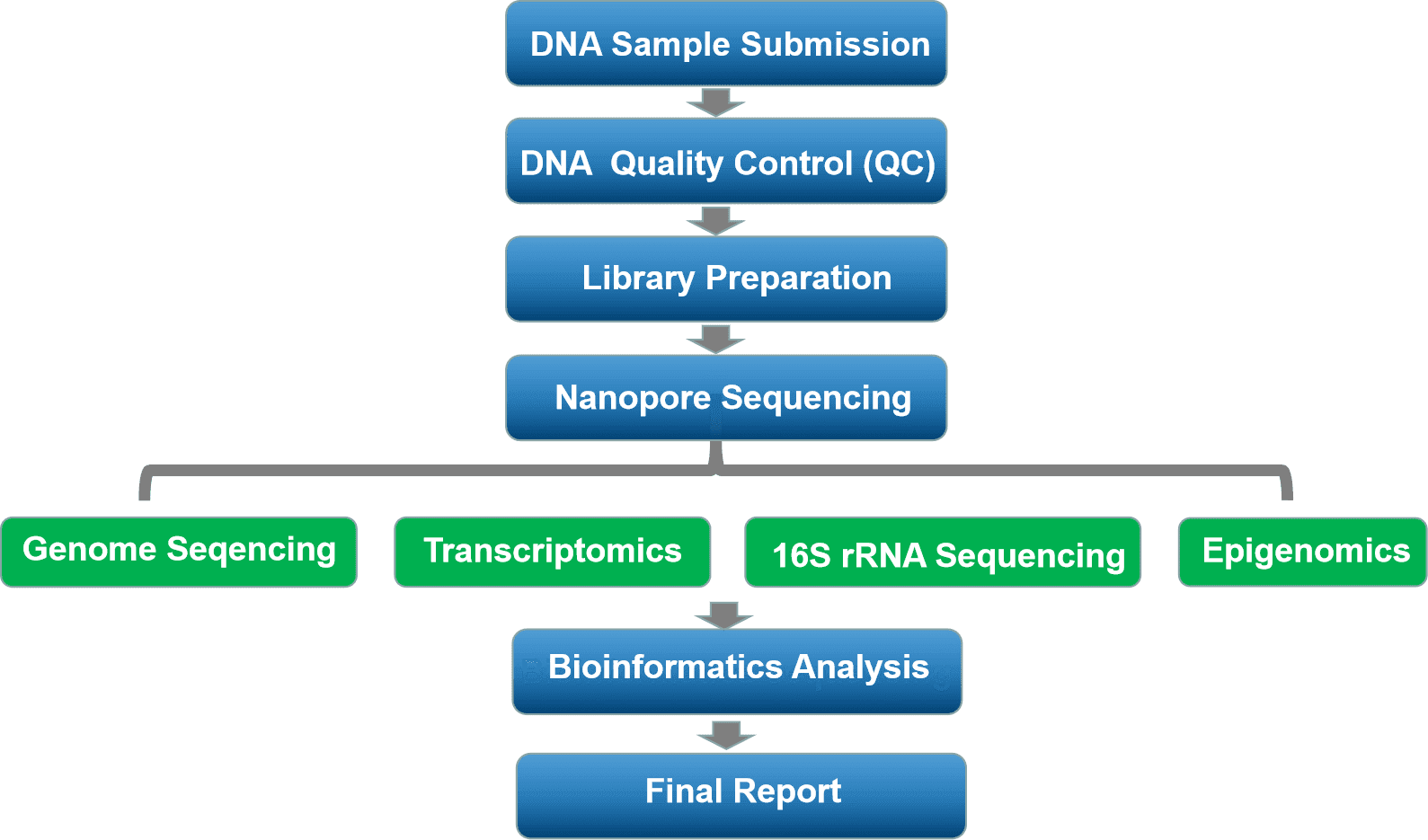
| Genome Analysis |
Genome assembly |
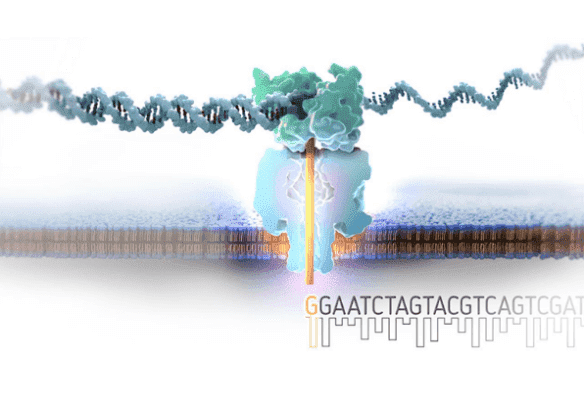
Determine the sequence of a genome using only randomly sampled sequence fragments |
| Mutation detection |
Detection of SNPs, indels, and CNVs |
| Structural variation |
Reveal mutational mechanisms and risk factors for chromosome rearrangement |
| Transcriptomics Analysis |
Gene isoform identification |
Detection, prediction, and characterization of RNA isoforms |
| Relative abundance of transcripts |
Estimation and visualization of the relative abundance of mRNA transcripts |
| Identification of AS events |
Identification and characterization of alternative splicing (AS) events |
| SNP calling |
Detection of single nucleotide polymorphisms (SNPs) |
| Expression analysis |
Detection of differentially expressed mRNA, lnRNA, circRNA, genes |
| 16S rRNA Sequencing Analysis |
Taxonomic assignment |
OUT clustering and filtering, taxonomic assignment |
| Phylogeny analysis |
Construct phylogenetic trees illustrating the evolutionary relationships among species |
| Statistical analysis |
PCA, Heatmap, VENN analysis, etc. |
| Epigenomics Analysis |
Data processing |
Exportation of sequence data and data processing such as adapter trimming |
| Alignment and assembly |
Genome assembly and polishing using NGS data |
| Methylation detection |
N6-methyladenine (m6A), 5-methylcytosine (m5C), and N4-methylcytosine (m4C) motifs, etc. |
| Custom Analysis |
More data mining upon your request |