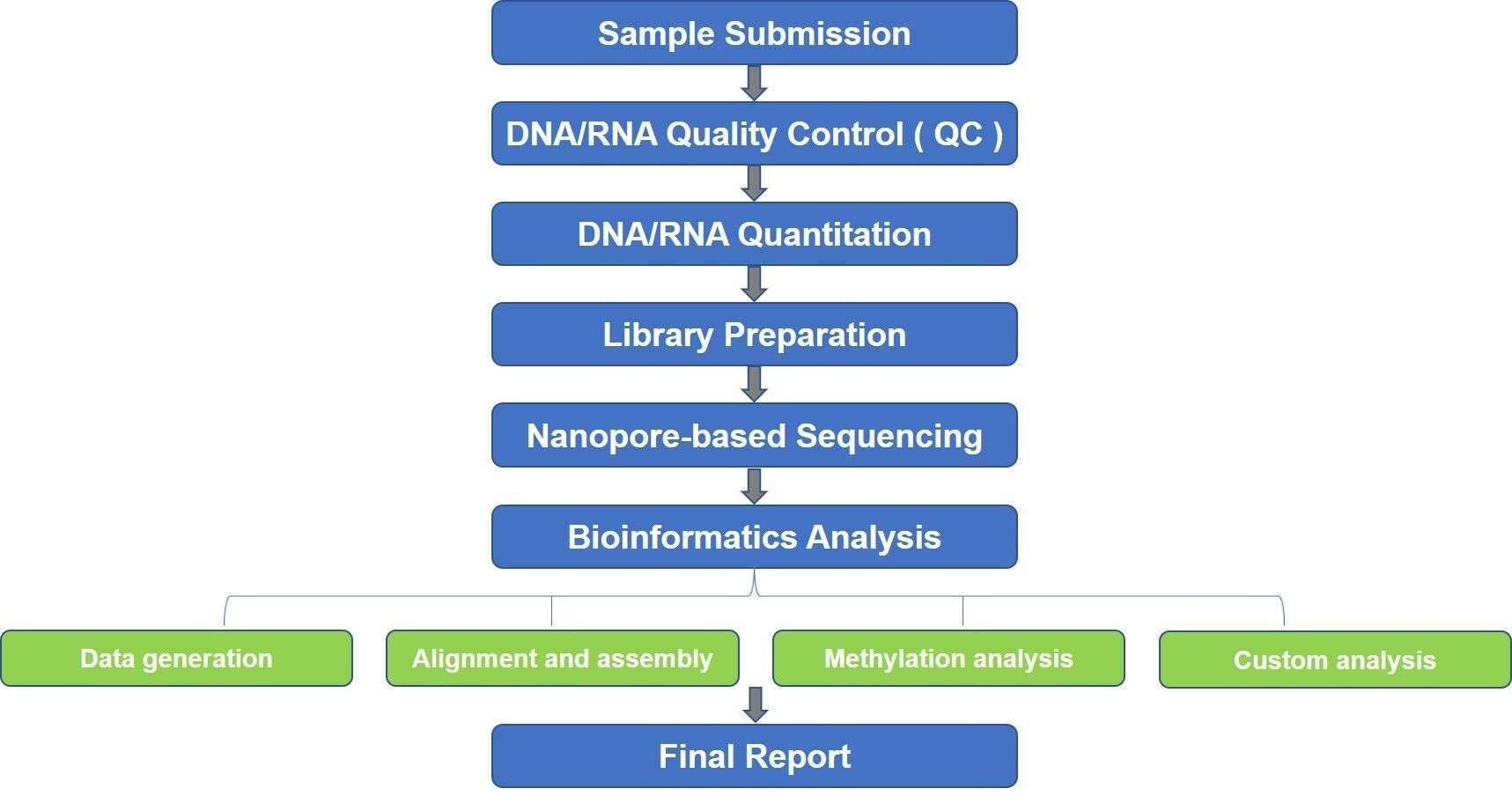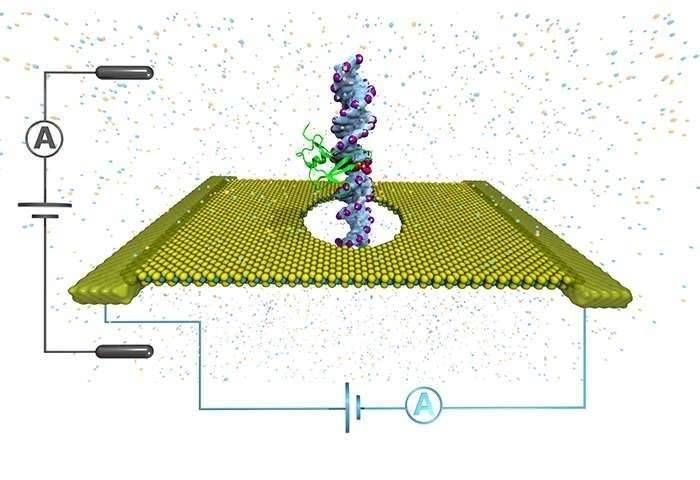
DNA is methylated by the MTase enzymes, which transfer a methyl group from S-adenosyl-l-methionine (SAM) to the appropriate positions of target bases. MTases function either alongside a cognate restriction enzyme as part of a RM (restriction-modification) system or as ‘orphans’ that lack a cognate restriction enzyme. Nanopore sequencing instruments rely on engineered biological nanopores embedded in a lipid membrane to sequence single-stranded DNA (ssDNA). A voltage is applied across the membrane, and ssDNA is ratcheted through a biological nanopore by a molecular motor protein bound to the DNA library molecule. The ionic current flowing through the nanopore depends on the set of nucleotides occupying the constriction point. The methylated nucleotides introduce distinct current patterns which can be detected. Nanopore-based epigenomics allows for simultaneous detection of DNA or RNA sequence, base modifications, and more comprehensive chromatin capture studies. We use the powerful metaepigenomics, a powerful approach, to identify a vastly unexplored microbial DNA methylation systems.
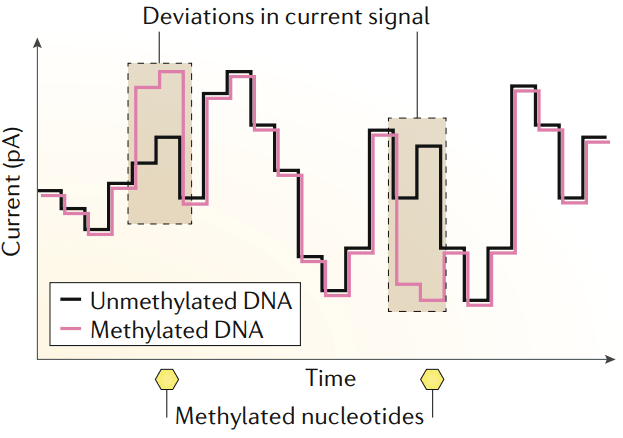
Compared to SMRT-based epigenomic sequencing, nanopore-based epigenomics analysis can detect more types of DNA modifications, such as N6-methyladenine (m6A), 5-methylcytosine (m5C), and N4-methylcytosine (m4C) motifs. The roles of methylation in regulating gene expression, virulence, and pathogen-host interactions can also be understood using this analysis. As sets of methylated motifs and MTases can vary widely, even between closely related microbial strains, nanopore-based epigenomics analysis is expected to enable differential methylation analyses between microbial populations.