What Is Microbial Diversity?
Microbes are tiny living organisms that are found in almost every habitat present in nature, including geological environments such as soil, deserts, oceans, and rocks. They also reside in our bodies such as in oral cavity, on the skin, and in gastrointestinal tract. The primary groups of microorganisms are bacteria, archaea, eukaryotes, and viruses. According to a new estimate, there are about one trillion species of microbes on earth, and still 99.999% of them have yet to be discovered, forming communities of astonishing diversity in their hosts or habitats.
Microbial diversity is a multi-latitude concept that includes:
- Microbial genetic diversity
- Microbial species diversity
- Microbial ecosystem diversity
Significance of Microbial Diversity Research
Microbes are the most abundant biodiversity resources on earth, and the development of microbial resources is fundamental not only for the resource exploitation and utilization but also for the maintenance and conservation of global genetic resources – discovery of new organisms and genes that are valuable to biotechnology, diversity patterns that can be used for the monitoring and prediction of environmental alternation, the role that microbiome plays in human health and homeostasis, and the development of novel genetic tools… The significance of microbial diversity research makes it one of the main driving forces for the development of life science in the 21st century.
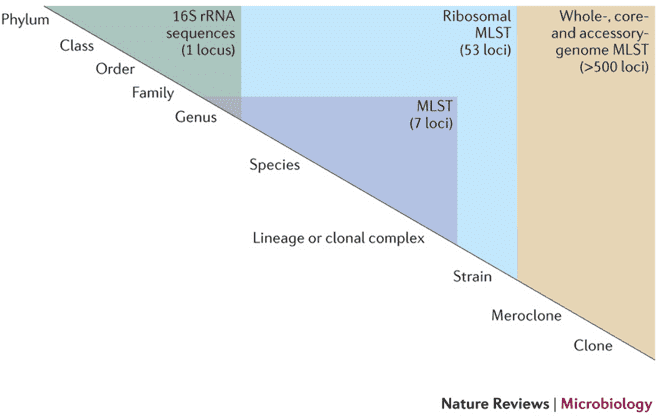 Figure 1: Discrimination level of different sequencing schemes (Maiden, 2013)
Figure 1: Discrimination level of different sequencing schemes (Maiden, 2013)
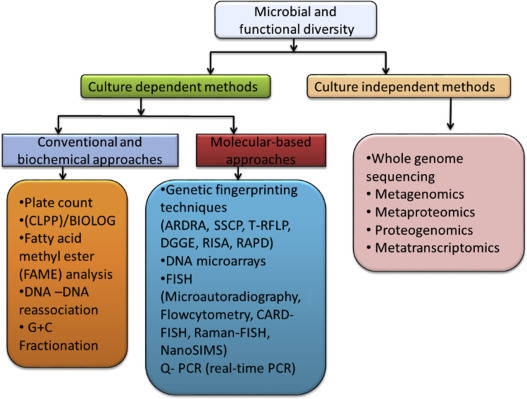
Diversity Research Techniques
Recent advancement in genetic technologies have promoted the soaring of microbial diversity research, allowed for high-throughput screening and analysis coupled with powerful bioinformatics platforms.

DNA fingerprinting: also known as DNA typing or DNA profiling, is a method used for DNA comparison or determination of an individual's DNA characteristics by studying the unique pattern of a series of DNA fragments after separating them by gel electrophoresis.

DNA microarray: also known as gene chip, in which a series of sequences or genes are orderly arranged in microarray to detect target DNAs or RNAs (most commonly as cDNA), which enables researchers to identify microbes and analyze the expression of thousands of genes in a single reaction.
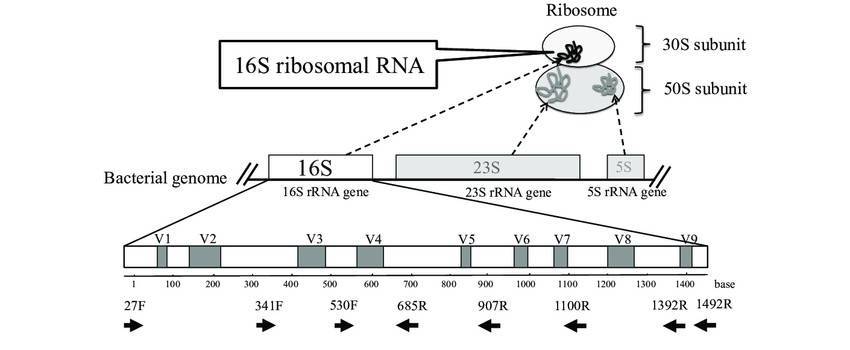
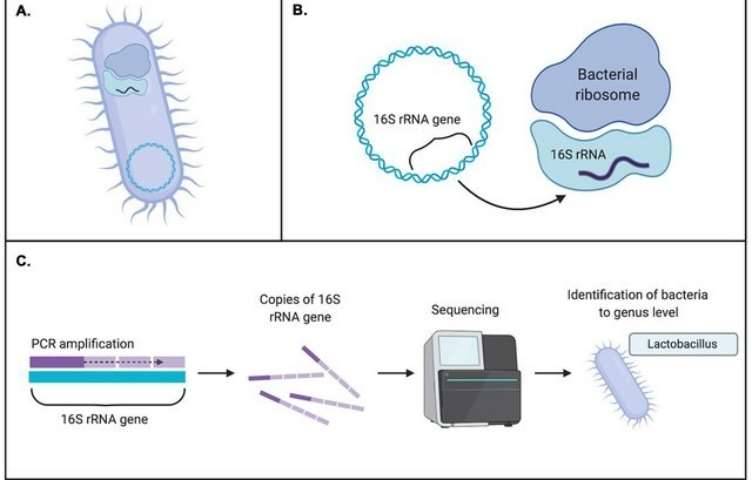
16S/18S/ITS sequencing: ribosomal RNA is the predominant form of RNA composed around 80% of cellular RNA, consisting of multiple conservative and variable regions. It can be used to identify and compare microbes through convenient amplification and distinguishable sequences. The most commonly sequenced fragments distribute within 16S, 18S and ITS.

Whole-genome sequencing (WGS): WGS works as a powerful tool to delivers a comprehensive view of the entire genome. It can be applied in multiple areas including diagnosis, tracking disease outbreaks, characterizing causative mutations and identifying drug-resistance genes. The major processes of whole-genome sequencing workflow are alignment, variant calling, annotation and calculating related metrics.

Metagenomic sequencing: metagenomic sequencing (or whole-genome shotgun sequencing, WGSS) read billions of DNA base pairs in a single run without selective enrichment of a specific population, is not only capable of profiling microbial community composition and diversity but also provides the information to reveal the functions encoded by the genomes.
Reference
- Maiden M. C.; et al. A nomenclature for discussing the genetic relatedness of bacteria. Nature Reviews Microbiology, 2013, 11: 728–736.