The gut microbiome is the overall composition of microorganisms, bacteria, viruses, protozoa, fungi, and their collective genetic material that live in the gastrointestinal tract (GIT). There are multiple organisms that can be seen in specified locations in the gastrointestinal tract (as seen in Figure 1) which allows them to have specific contributions to metabolism and absorption of nutrients can be seen in specified locations. The human gut microbiota is partitioned into numerous gatherings called phyla. The gut microbiota is involved fundamentally in four primary phyla which incorporate Firmicutes, Bacteroidetes, Actinobacteria, and Proteobacteria. While microbes colonize the human body, including oral holes, placenta, vagina, skin, and GIT, most microscopic organisms dwelling inside the GIT, with most of the dominatingly anaerobic microorganisms housed in the colon.
The development and modification of the gut microbiome are influenced by an assortment of variables including birthing and feeding method, exposure to stress, climate, diet, prescriptions, phase of the lifecycle, and comorbid infections. The arrangement of every microorganism present in the GIT has its specification and assumes a function in orchestrating helpful naturally dynamic particles, regulating insusceptible reactions, conduct, and disposition.
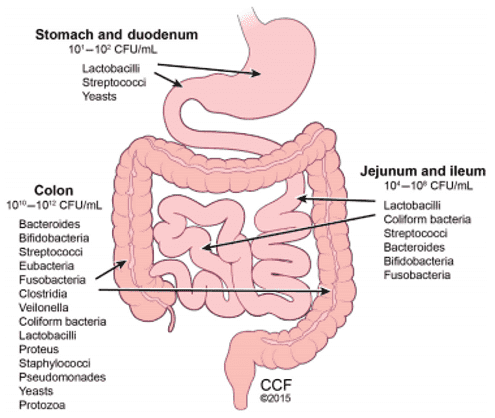 Figure 1. Gut microbiota predominance. (Cresci, 2019)
Figure 1. Gut microbiota predominance. (Cresci, 2019)
Ways to Improve Gut Microbiota
Many different medical treatments have been used to improve the gut microbiota and one of the common techniques is the fecal microbiota transplantation (FMT). This microbiome treatment has been set up as a powerful treatment for repetitive Clostridioides difficile disease and might be a successful methodology for decreasing intestinal ARO colonization and by reestablishing microbial network organization and capacity. Prebiotics, nonliving nondigestible compounds from fiber or carbohydrates have been shown to increase levels of beneficial bacteria, playing a role in the promotion of our health. Prebiotics also is said to improve the multiplication of helpful microorganisms or probiotics, to boost practical changes in the human microbiome. Besides dietary fibers and FMT, live active microorganisms called probiotics are said to have beneficial effects to its host when administered in adequate amounts. Probiotics may reestablish the piece of the gut microbiome and acquaint valuable capacities with gut microbial networks, bringing about enhancement or anticipation of gut aggravation and other intestinal or foundational ailment phenotypes. Probiotics can likewise antimicrobial operators or metabolic intensifies that stifle the development of different microorganisms, or then again contest for receptors and restricting locales with other intestinal organisms on the intestinal mucosa. Probiotics can affect our gut microbiota by implication too, for instance by impacting our insusceptible framework which thusly impacts our colonizing microbiota.
Techniques Used to Study Gut Microbiome
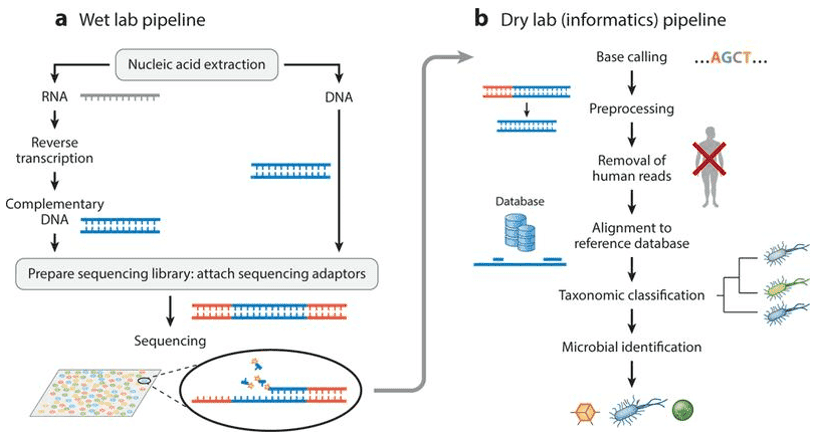
Gut microbiomes can be studied in many different techniques. Different organisms in the gastrointestinal tract have different and unique genomes in their structure. With these differences, the approach of genomics can be used to understand the capabilities of the different organisms within its community, its relationship with its members, and the environment.
Using the transcriptome or the complete set of all the RNA molecules expressed in the organism, gut microbiomes can be studied furthermore through the means of transcriptomics. The proteins present in the microorganisms that live in the GIT can then be also analyzed for studying the classification and function of the microbiota through the use of proteomics wherein large sets of proteins are being fractionated, gone under mass spectrometry, and analyzed. Furthermore, the metabolites produced by the microorganisms present in the GIT can be taken advantage of and be used for analyzing and studying the importance of the microbiota in the GIT. This is through the means of metabolomics where small molecules undergo nuclear magnetic resonance, mass spectrometry, or gas chromatography-mass spectrometry in order to identify the phenotypes of the microorganism.
Identifying Gut Microbiome Composition
One of the ways microorganisms in the GIT can be obtained through stool sampling. In many investigations, the bacterial constituents of a microbial population found in the stool sample are recognized by the technique known as the 16S rRNA gene sequencing. Targeting the 16s rRNA gene would lead to the identification of these microorganisms. A specified gene which is the 16S rRNA gene is isolated and then amplified through polymerase chain reaction.
References
- Collado M., Meriluoto J., Salminen S. Role of commercial probiotic strains against human pathogen adhesion to intestinal mucus. Lett Appl Microbiol. 2007, 45: 454-460.
- Cresci, G. A. M., Izzo, K. Adult Short Bowel Syndrome, Gut Microbiome. 2009, 45–54.