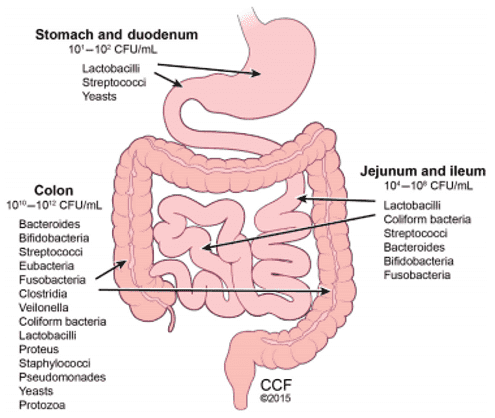
Our bodies have a mutually beneficial relationship with trillions of microbes currently living within the gastrointestinal tract. The microbiome is known simply as numerous species creating this population. Through improving digestion, enforcing metabolism, convening resistance to pathogens, and modifying host immunity, host-microbiome interactions can have abrupt effects on health. In order to prepare the immune system and ensure immune homeostasis, balanced microbial communities can give positive. Dysbiosis is a shift in the normal component of the microbiome that can start chronic inflammation, infractions of the epithelial barrier, and detrimental bacteria infestation. Carcinogenesis has been correlated with all these variables.
Traditionally, by culture and subsequent staining, representatives of microbial communities were defined, which prevented the investigation of non-culturable species and strains. However, the research of the microbiome has been pioneered by next-generation sequencing (NGS) techniques. NGS allows for continuous analysis of numerous species within a bacterial diversity without the need to culture individual organisms. It has become necessary to gauge species variation and evaluate dynamic fluctuations in bacterial diversity with the growth of bioinformatics tools to analyze large volumes of new knowledge. As institutions began to realize the important role that microbes participate in health and disease, new initiatives to help the research community were initiated. The Human Microbiome Project (HMP) financed by the NIH and the European Metagenomics of the Human Intestinal Tract (MetaHIT) both utilize NGS-based data to determine productive human microbiome reference genome datasets.
Multiple microbial species have been engaged in supporting local inflammation-associated tumor growth. More recently, microbes that affect the effectiveness of anticancer drugs have been said to have systemic effects on the host. Even though the processes of these effects are not clearly illustrated, immune system modulation is often a component. NGS' capability to evaluate complex microbial communities may assist to identify new host-microbe interaction methods that promote cancer or enhance the efficacy of drugs. This application note exemplifies several key findings on the impact of the microbiome on cancer progression and evaluations the technologies that can be applied to further evaluate relationships between host and microbe.
Microbial Sequencing Methods
NGS enables scientists to sequence numerous organisms in parallel, unlike capillary sequencing or PCR-based methods. NGS-based metagenomic sequencing can identify low-abundance members of the microbial community that may be overlooked or is too costly to recognize using other methods with the way to gain high sequence coverage per specimen. For numerous NGS techniques, Illumina offers simplified frameworks that can give a critical genetic perspective into archiving microbiome species and tracking their complexities.
For microbiome researches, three fundamental NGS applications are accessible:
2) Metagenomic shotgun sequencing
3) Assessment of the metatranscriptome.
Accurate improvement through each sequencing interface is made possible by high-quality materials and user-friendly applications. The selection of the system depends on the subject being asked about the study.
References
- Zama D, Biagi E, Masetti R, et al. Gut microbiota and hematopoietic stem cell transplantation: where do we stand?. Bone marrow transplantation. 2017 Jan;52(1).
- Lee YK, Mazmanian SK. Has the microbiota played a critical role in the evolution of the adaptive immune system?. science. 2010 Dec 24;330(6012).