MicroSEQ Identification by 16S/D2 Region Sequencing: Introduction, Protocol, and Applications
Inquiry >Introduction to MicroSEQ
For microbial identification and stratification, ribosomal RNA (rRNA) sequencing has been observed to have the highest accuracy. Applied Biosystems MicroSEQ system, which uses 16S rRNA (for bacteria) and LSU-D2 region (for fungi) sequencing to identify microbes, is a highly accurate and fast method for microbial identification. Microbial identification down to the genus or species level has a rate of success of over 99 percent.
MicroSEQ-based microbial identification is much faster and more accurate than presently used phenotypic techniques. Sequence analysis of the 16S rRNA gene and the LSU-D2 region can be utilized to identify badly described, rarely isolated, or phenotypic variant strains and is routinely used to identify mycobacteria. This technique can be used to find new pathogens and bacteria that haven't been cultured before. For pharmaceutical companies, biotech corporations, medical device corporations, public health laboratories, and government agencies, the MicroSEQ system has been used for microbial identification duties such as routine QC microbiology.
MicroSEQ Verification Processes
Laboratory authentication, bioinformatic authentication, and phylogenetic authentication are the three authentication processes used by the MicroSEQ.
The sequence acquired from the sample using MicroSEQ ID procedures, as well as the sequence-verified for quality, are included in the laboratory authentication. Bioinformatic authentication, on the other hand, entails the description of sequences that have been verified against public databases such as NCBI and sequences that have been checked against existing MicroSEQ libraries. Finally, phylogenetic verification entails confirming specimen identification through phylogenetic analyses, as ascertained by sequence-based clustering of the sample with the assumed group of organisms.
Applications
Identification of Bacteria
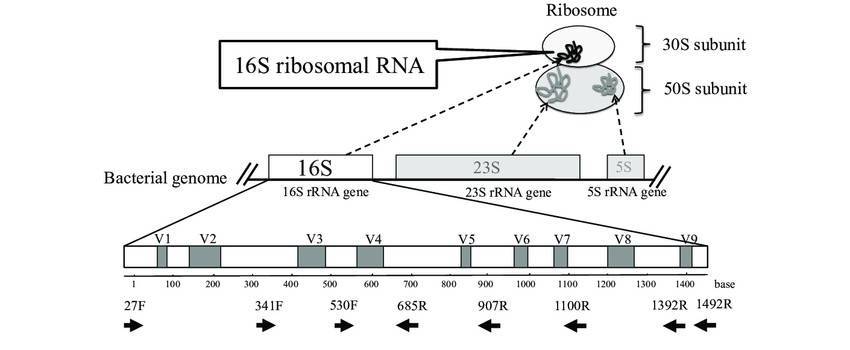
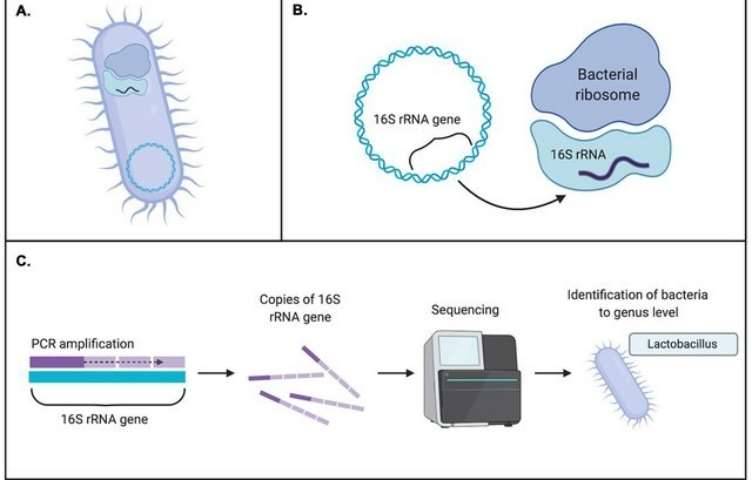
The 16S ribosomal RNA (rRNA) gene sequence is the target for bacterial identification (rDNA). The 16S rRNA is a ubiquitous component of the small subunit of a prokaryotic ribosome and can therefore be utilized to examine phylogenetic relationships among all bacteria.
Users can sequence either the first 500 bp or the entire 1,500 bp of the 16S rDNA gene using the MicroSEQ® ID technique. For both choices, kits and the supporting verified library databases are accessible. For routine identifications, the first 500 base pairs are sufficient to cover three of the 16S gene's nine hypervariable areas.
In some cases, the 500-bp region is insufficient to distinguish between closely related bacteria, necessitating a more comprehensive full-genome read (taking into consideration all the hypervariable areas). Furthermore, when describing a new species, the entire 1,500-bp sequence must be sequenced.
Identification of Fungi and Yeast
Both the expansion area D2 of the bigger rRNA molecule in the large subunit of the eukaryotic ribosome (LSU-D2) and the ITS areas have been effectively used for the identification and classification of fungi down to the species level using comparative sequence analysis. When used for sequence-based fungal identifications, the ITS region has more variations, which can result in many more "sequence types" than the D2 region. Because fungal genomes can have up to 100 copies of the rDNA cluster, it's important to know how to distinguish between simplified sequence variability and sequence variability that represents actual relatedness. The MicroSEQ® ID fungal database is based on sequences from LSU-D2 rDNA, and it aids in the identification and classification of fungal species in routine identification assessments.
References
- Cloud JL, Conville PS, Croft A, Harmsen D, Witebsky FG, Carroll KC. Evaluation of partial 16S ribosomal DNA sequencing for identification of Nocardia species by using the MicroSeq 500 system with an expanded database. Journal of clinical microbiology. 2004 Feb;42(2).
- Woo PC, Ng KH, Lau SK, et al. Usefulness of the MicroSeq 500 16S ribosomal DNA-based bacterial identification system for identification of clinically significant bacterial isolates with ambiguous biochemical profiles. Journal of clinical microbiology. 2003 May;41(5).