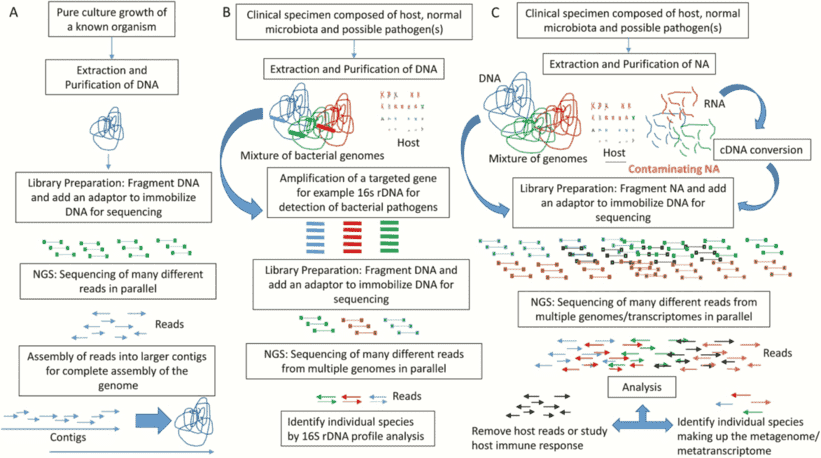
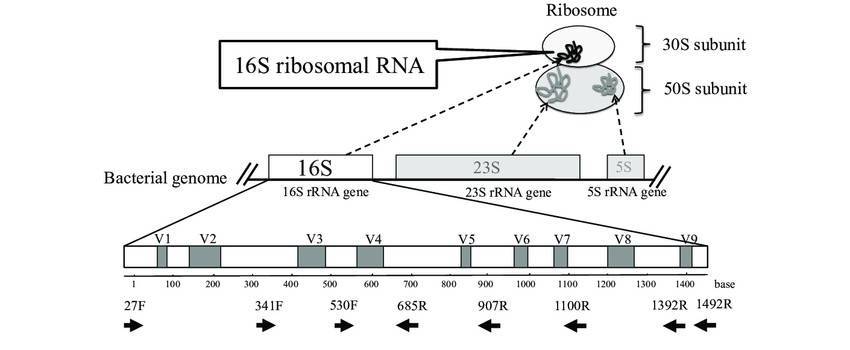
Small subunit ribosomal RNA (especially 16S rRNA) is presently the bacterial molecular systematics molecule of preference, and several dedicated databases promote this large body of information. Some protein-coding genes, however, may have selective benefits over rRNA data. Higher sequence variation degrees allow distinction of closely related strains, while phylogenetic analysis of distantly related strains and more precise sequence alignment allows the potential to interpret DNA to protein sequences. In order to supply a focal point for bacterial recognition and characterization using protein-coding genes, the Identification and Classification of Bacteria (ICB) database was created. The database focuses on the provision of immense sequence data support resources, initially focusing specifically on DNA gyrase subunit B (gyrB), a type II topoisomerase seen in bacteria that can introduce negative supercoils into a loosen, enclosed circular DNA molecule.
Numerous publications have proposed that gyrB is an appropriate gene for bacterial phylogeny, having key qualities such as limited horizontal transmission and involvement in all bacterial groups since 1995 when universal primers for the gene became accessible. A significant quantity of gyrB sequence data has been produced over the last 5 years and the ICB database seeks to achieve this data accessible to the scientific community in a feasible layout.
Significance of gyrB on Bacterial Research
In many categories in other phyla, the gyrB gene is said to be an effective marker for phylogenetic study. The phylogenetic analysis focused on the patterns of the gyrB gene has been seen in some experiments to be compatible with DDH and phenotypic comparison. To evaluate the phylogenetic relations of marine isolates within the phylum Bacteroidetes, gyrB gene sequencing was adapted and involved two species of Flavobacterium. Moreover, within the framework of genome projects, more gyrB sequences from Flavobacterium species are becoming obtainable.
Several Flavobacterium strains were segregated in a previous study of aquatic and terrestrial microbial mats in Antarctica, which displayed a low resemblance to the Flavobacterium species, based on the partial or full 16S rRNA gene sequence. In a study, to evaluate the variety of isolates in more detail and to clarify the usefulness of gyrB as a phylogenetic marker for phylogeny in the Flavobacterium genus, the gyrB gene sequence of 33 of these new Antarctic isolates and of the type strains of related Flavobacterium species was defined. It also compared the phylogeny based on the near-complete sequences of the 16S rRNA gene.
A Case Study Focused On gyrB
In a study, using the ClustalW program, the available gyrB nucleotide sequences of Yersinia spp were coordinated and conserved regions were utilized to construct Yersinia consensus primer sets. Amplification of the PCR was performed in a 25 ml reaction mixture containing 20 pmol per primer, gyrF (50-GTTTCC GGCGTTTGCACGG-30), and gyrRRCACG-30 respectively (50 -GTGATAACGCA ATTTGTCCGGG-30). The amplification was carried out with the following temperature profile: initial denaturation for 5 min at 95 C, followed by 30 cycles of 95 C for 1 min each, 62 C for 1 min and 72 C for 1 min, and final extension for 10 min at 72 C. PCR products have been purified and sequenced. The polymorphism in the 1143 bp gyrB amplicons was assessed by restriction digestion using AluI, HinfI, and MspI enzymes. On 2.5 percent agarose gels in a 1X TBE buffer, restriction fragments were segregated.
References
- Schoeffler AJ, May AP, Berger JM. A domain insertion in Escherichia coli GyrB adopts a novel fold that plays a critical role in gyrase function. Nucleic acids research. 2010, 38(21).
- Gulati PS, Virdi JS. The rrn locus and gyrB genotyping confirm the existence of two clonal groups in strains of Yersinia enterocolitica subspecies palearctica biovar 1A. Research in microbiology. 2007, 158(3).