Microorganisms are the most abundant living organisms in the world. They are the dominant drivers of biogeochemical processes in nature. The ocean is a vast uncharted territory when it comes to understanding microbial diversity, community structure, and ecological determinants. One of the interesting facts about oceans is that it is one of the most dilute ecosystems in terms of nutrient availability but still, many species of bacteria, archaea, viruses, plankton, and other microorganisms lie beneath the surface of the world's oceans waiting to be identified and classified. The study of ocean microbiome is a relatively new field of oceanography that started only about a few decades ago. Up until now, there is much to learn about the influence of these microorganisms on the biological processes underwater.
Most marine microbial communities are composed of a few dominant species and several rare species, depending on the wideness of the ocean, availability of resources, depth, and structure, and composition of organic matter. This information helps researchers to predict and prepare model communities for a particular ocean ecosystem for a particular environmental condition. DNA sequencing technologies such as metagenomic sequencing and DNA barcoding has been useful in studying the ocean microbiome. 16S ribosomal RNA gene sequencing was first used in surveying natural environments. Later on, other meta-omics technologies have also been used to improve the microbial inventory of the ocean.
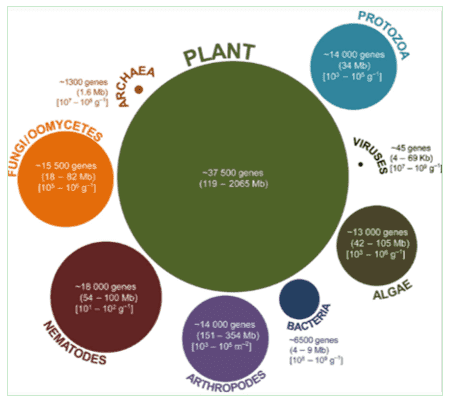
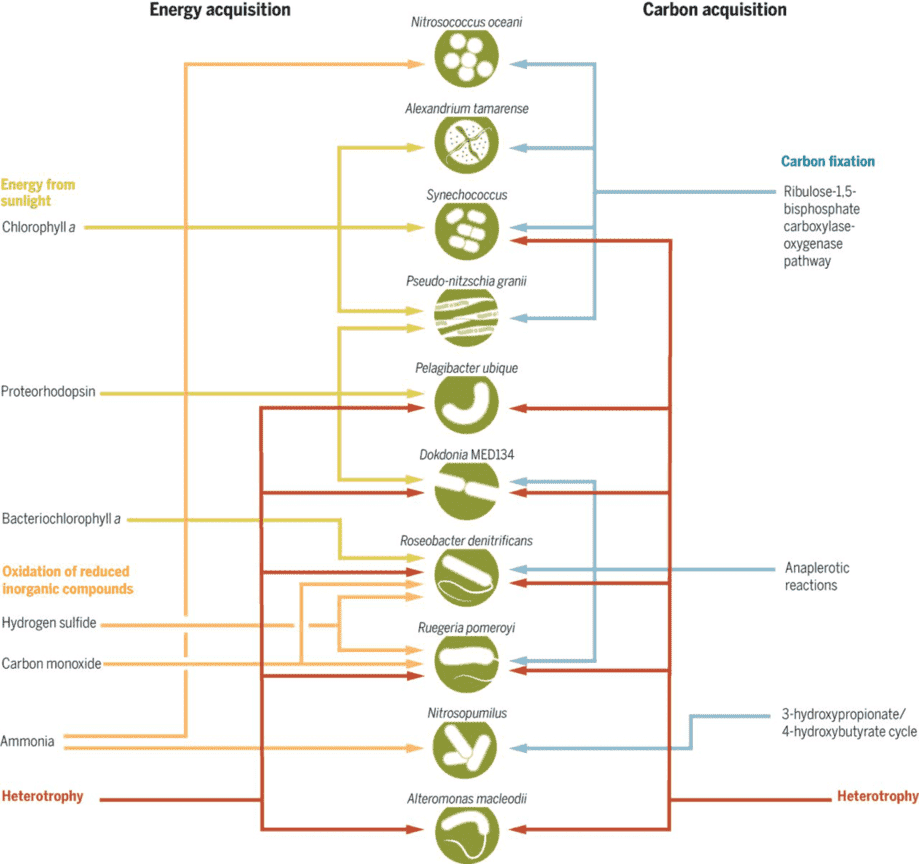 Figure 1. Trophic categories of the ocean microbiome. (Moran, 2015)
Figure 1. Trophic categories of the ocean microbiome. (Moran, 2015)
Through the years, several distinct and recurring bacterial communities have been discovered, including their relative abundance and patterns of occurrence, by means of 16S ribosomal RNA sequence analysis. The taxonomic inventories of the world's ocean microbiome continue to grow as scientists are able to provide more characterization and profiling of marine microbes and mapping the seasonality of microbial populations. Some studies have also been dedicated to the understanding of the involvement of microorganisms in ocean biogeochemical processes, and the production of ocean bioproducts such as vitamins, hormones, and antibiotics.
The world's ocean has a population of an estimated 104 to 106 cells per milliliter and contains immense genetic diversity. With the development of metagenomics technologies and computational analysis, the study of taxonomic and genomic characteristics of ocean microbiomes has been more accessible to researchers and can help in designing predictive models for species abundance, interspecies interaction, community connectivity, and the effects of environmental conditions to these microbial communities.
In 2015, a group of oceanographers and researchers attempted to map a larger coverage of the structure and function of the world's ocean microbiome. Using metagenomic sequencing technologies, they were able to analyze more than 7 tera bases worth of metagene from 68 locations in the world. They were able to identify over 40 million species of microorganisms including viruses, prokaryotes, and picoeukaryotes. One of the most significant findings in the research was that there is more than 70% similarity in species abundance between the ocean microbiome and the human gut microbiome.
The ocean microbiome holds a lot more information about bacteria, virus, and other picoeukaryotes, their interactions, and how they influence the flux of materials in the ocean. With the help of sequencing technologies and computational analysis, we are to discover mechanisms of interaction that may also be applied to other microbiomes in an attempt to better understand microbial communities.
References
- Moran MA. The global ocean microbiome. Science. 2015, 350(6266).
- Sunagawa S, Coelho LP, Chaffron S, et al. Structure and function of the global ocean microbiome. Science. 2015, 348(6237).