The introduction of next-generation sequencing (NGS) has provided us with a broader and deeper perspective of genetic research. While whole‐genome sequencing may capture all possible mutations, targeted NGS that focuses on specific regions of the genome remains a widely-used, efficient, and cost-effective strategy, which offers a focused approach of the most phenotype‐altering regions for NGS. The enrichment is a critical step of target NGS. It is the selection of the regions of interest out of the whole genome, allowing researchers to choose to only study a fraction of the genome where specific genes, exons, or other genomic regions of interest present. Moreover, targeted NGS simplifies downstream bioinformatics analysis, provides greater sequencing depth, and increases sequencing sensitivity, which is valuable in variant calling and gene expression studies. Two fundamental target enrichment strategies can be employed prior to NGS – capture and amplification.
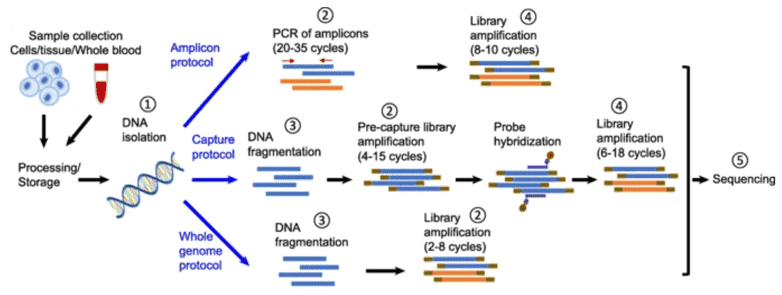 Figure 1. Illustration of the major protocols of typical next-generation sequencing. (Ma 2019)
Figure 1. Illustration of the major protocols of typical next-generation sequencing. (Ma 2019)
Hybridization-Based Target Enrichment
The "capture" can be realized through the hybridization of fragmented DNA with baits complementary to regions of interest, either on a microarray chip or free in solution. In-solution hybridization capture can be scalable and easily automatable, and rules out the shortcomings of microarray hybridization such as high cost, limitations to the number of samples and large sample requirements. Thus, solution-based hybridization becomes the mainstream approach of capture-based enrichment for NGS.
In a hybridization assay, the protocols start with random shearing of the genes, denatured by heating. Randomly sheared overlapping fragments are captured by DNA or RNA single-stranded oligonucleotides specific to the region of interest. DNA bound to the baits is separated from background DNA, the baits are removed and DNA enriched for regions of interest is taken forward to the second round of PCR, followed by NGS of the enriched nucleic acids.
Amplification-Based Target Enrichment
Pre-NGS targeted enrichment can also be carried out using the PCR strategy, which is known as the amplicon-based target enrichment. Embarked with greater primer-probe specificity compared to hybrid-based capture, higher on-target sequencing reads can be obtained by amplicon-based enrichment, which has been preferentially used on the basis of short preparation time and small DNA input amounts. Thus, amplicon-based NGS enables fast, accurate, and cost-effective detection of genes of interest.
In an amplicon-based target enrichment assay, genomic DNA is firstly fragmented by enzymatic or mechanical methods. The following step involves the amplification of the regions of interest using sequence-specific primers, where regions of interest could vary greatly among exome, rRNA, miRNA, etc. In this step, PCR could be single or multiplex, generating single or multiple amplicons. Then amplicons representing each region of interest are pooled to create a library of fragments for sequencing.
A Comparison of Hybridization and PCR Strategies
| Feature | Hybridization Capture | Amplicon Sequencing |
|---|---|---|
| Input Amount | >1 μg DNA | 10–100 ng |
| Number of Targets | Virtually unlimited | Limited |
| Sensitivity | Down to 1% | Down to 5% |
| Workflow | Laborious and complex | Simple and fast |
| Cost | Varies | Generally lower cost per sample |
| Best-Suited Applications |
|
|
Generally speaking, amplicon-based approaches may be preferable for their simplified workflow and smaller amounts of required DNA. However, hybridization-based strategies are less likely to miss mutations and also perform better with respect to sequencing complexity and uniformity of coverage. There are also differences with regard to sample requirements, sensitivity, workflow, cost, etc. The choice of enrichment approaches for targeted sequencing is an important consideration, where all the above factors worth careful consideration.
References
- Ma X; et al. Analysis of error profiles in deep next-generation sequencing data. Genome biology. 2019, 20(1): 50.
- Gasc C; et al. Sequence capture by hybridization to explore modern and ancient genomic diversity in model and nonmodel organisms. Nucleic acids research. 2016. 44(10): 4504-18.
- Samorodnitsky E; et al. Evaluation of hybridization capture versus amplicon‐based methods for whole‐exome sequencing. Human mutation. 2015, 36(9): 903-14.