Metagenomic Sequencing Data Analysis Service: Unlocking Insights into Microbial Communities
Introduction
Metagenomics, the study of genomes within microbial communities, has revolutionized our understanding of the microbial world. Metagenomic sequencing plays a crucial role in unraveling the complexities of these communities, enabling us to explore species composition, gene function, and metabolic pathways associated with specific environments. CD Genomics, a leading company in the field of genomics and bioinformatics, offers comprehensive metagenomic sequencing data analysis services that empower researchers to gain deep insights into microbial communities.
Metagenomics Analyses Applications
Metagenomics is a molecular tool used to analyze DNA obtained from environmental samples to study the microbial communities present without obtaining pure cultures. Functional metagenomics allows high-resolution genome analysis of non-culturable microorganisms and correlates genomes with specific functions in the environment, which has diverse applications across various fields, ranging from environmental microbiology to human health.
Environmental Microbiology: Metagenomics is widely used to study the biodiversity, functional potential and ecological role of microbial communities in different environments such as soil, marine and freshwater ecosystems, which contributes to the understanding of microbial interactions, nutrient cycling and environmental factors Effects on the microbial community.
Bioremediation and Environmental Monitoring: Metagenomics is vital in bioremediation and environmental monitoring, which utilize microorganisms to break down pollutants. Researchers discern microbial genes and pathways related to pollutant degradation by scrutinizing contaminated sites' metagenomes. This assists in crafting efficient bioremediation approaches. Additionally, metagenomic analyses help identify and evaluate potentially harmful microorganisms in environmental samples, aiding in environmental monitoring.
Industrial Applications: Metagenomics is valuable in various industrial sectors, such as agriculture, food production, and biofuel development. By analyzing microbial communities in soil or plant samples, researchers can identify beneficial microbes that enhance plant growth, protect against pathogens, or contribute to nutrient cycling. In the food industry, metagenomic analyses assist in monitoring food safety, identifying spoilage microorganisms, and optimizing fermentation processes. Additionally, metagenomics aids in exploring microbial diversity for the production of enzymes, bioactive compounds, and biofuels.
Human Microbiome Research: Metagenomics has revolutionized our understanding of the human microbiome—the collection of microorganisms residing in and on the human body. By analyzing metagenomic data from various body sites, such as the gut, skin, and oral cavity, researchers can explore the composition, diversity, and functional potential of the human microbiome. Metagenomic analyses help in investigating the roles of the microbiome in health and disease, identifying potential microbial biomarkers, and understanding the impact of factors like diet, lifestyle, and medication on the human microbiome.
Infectious Disease Surveillance: Metagenomic analyses have emerged as a powerful tool in infectious disease surveillance, particularly for detecting and characterizing novel or emerging pathogens. By sequencing the genetic material in clinical samples, such as blood or respiratory specimens, researchers can identify infectious agents, including viruses, bacteria, and fungi. Metagenomics enables rapid identification and characterization of pathogens without the need for culture-based methods, facilitating early detection and response to infectious disease outbreaks.
Antimicrobial Resistance Studies: Metagenomics plays a significant role in comprehending the mechanisms and patterns of antimicrobial resistance (AMR) in Antimicrobial Resistance Studies. Researchers can identify AMR genes, determine their frequency, and examine the dissemination of resistance determinants by scrutinizing the metagenomic data from various sources such as environmental samples, animal microbiomes, or human populations. This type of analytical approach aids in monitoring the transmission of AMR among different reservoirs, identifying the possible origins of resistance genes, and devising solutions to tackle the worldwide issue of AMR.
Metagenomics Data Analysis Workflows
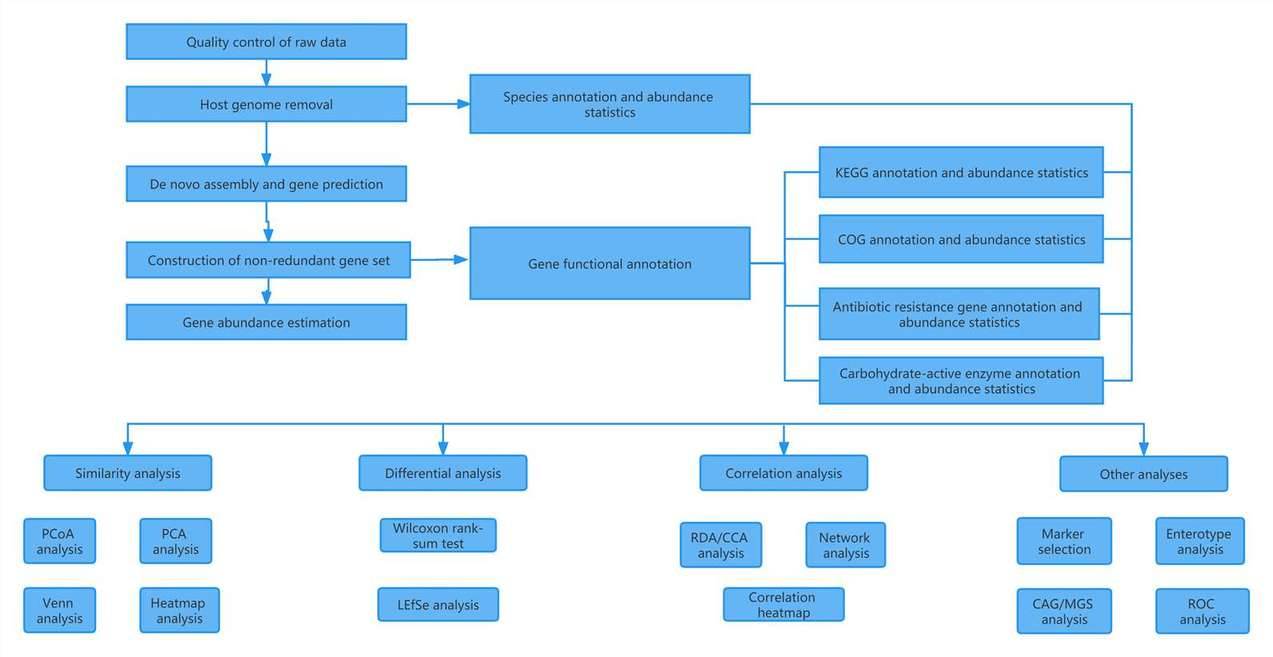
Metagenomic sequencing data analysis involves a series of computational steps to extract meaningful information from raw sequencing data. CD Genomics employs robust workflows to ensure accurate and comprehensive analysis of metagenomic data. The following steps are typically included in metagenomics data analysis workflows:
Pre-processing: Raw high-throughput sequencing data undergo quality control, including read trimming and removal of low-quality reads and sequencing adapters.
Taxonomic Profiling: Species and strain composition profiling is performed using specialized algorithms to assign reads to specific taxonomic groups. This information provides insights into the diversity and abundance of microbial communities.
Functional Annotation: Genomic components are analyzed to identify genes and their functions. This step involves assigning functional annotations to genes, enabling researchers to understand the potential metabolic capabilities of the microbial community.
Comparative Analysis: Comparative analysis among samples allows for the identification of differences in species composition, gene content, and functional pathways. This analysis provides valuable insights into the variations and dynamics of microbial communities across different environments or conditions.
Visualization: CD Genomics' metagenomic data analysis services include the generation of visualizations to present the results effectively. These visualizations help researchers interpret and communicate the complex data in a comprehensible manner.
Metagenomic Data Analysis Service
CD Genomics' metagenomic data analysis services encompass a wide range of applications and features to meet the diverse needs of researchers. The following are the key components of CD Genomics' metagenomic data analysis services:
Metagenomic Data Analysis
CD Genomics provides comprehensive metagenomic data analysis, focusing on species composition and abundance analysis, gene composition and function analysis, as well as metabolic pathway analysis associated with specific environments. Through state-of-the-art bioinformatic tools and pipelines, CD Genomics enables researchers to gain a deep understanding of microbial communities and their functional potential.
Metatranscriptomic Data Analysis
In addition to metagenomic data analysis, CD Genomics offers metatranscriptomic data analysis services. This advanced approach allows researchers to study gene expression within microbial communities. Metatranscriptomic analysis provides insights into the transcriptional activities and functional dynamics of microbial communities. By analyzing the RNA transcripts present in a sample, researchers can identify actively expressed genes, regulatory pathways, and responses to environmental conditions.
CD Genomics' metatranscriptomic data analysis services include functional profiling to understand the gene expression patterns and activities within the microbial community. This analysis enables researchers to explore the functional roles of specific genes and pathways in various biological processes.
Advanced Joint Metagenomic and Metatranscriptomic Data Analysis
CD Genomics specializes in advanced joint metagenomic and metatranscriptomic data analysis, which integrates both types of data to provide a more comprehensive understanding of microbial communities. This approach enables researchers to correlate gene expression with the presence and abundance of specific microbial species or strains. By combining metagenomic and metatranscriptomic data, CD Genomics facilitates gene-level, species-level, and strain-level analysis, leading to more accurate insights into the functional potential and activities of the microbial community.
Network Studies in Microbiome and Host-Microbiota Interactions
The interactions between microbial communities and their host organisms play a crucial role in various biological processes and human health. CD Genomics offers network analysis services to study microbiome and host-microbiota interactions based on high-throughput multi-omics data. By integrating metagenomic, metatranscriptomic, and other omics data, researchers can construct complex networks that reveal the relationships between microbial species, host factors, and environmental variables. These network studies provide valuable insights into the functional connections and dependencies within the microbiome and their impacts on host health and disease.
Causality Study between Human Microbiota and Diseases
Understanding the role of the human microbiota in health and disease is a rapidly evolving field of research. CD Genomics provides specialized services for causality studies, aiming to unravel the causal relationships between human microbiota and various diseases. Through integrative analysis of metagenomic and clinical data, researchers can identify microbial signatures associated with specific diseases or conditions. This analysis helps in elucidating the potential mechanisms through which the microbiota influences disease development, progression, and treatment response.
Existing Tools/Pipelines Recommendation, Analysis Training, and Guidance
CD Genomics offers expertise and guidance in selecting the most suitable bioinformatic tools and pipelines for metagenomic data analysis. With years of experience in the field, CD Genomics stays updated with the latest advancements and best practices in bioinformatics. The company provides recommendations on existing tools and pipelines that align with the specific research objectives and data requirements.
Additionally, CD Genomics offers training and guidance to researchers in metagenomic data analysis. The expert team provides valuable insights, practical tips, and hands-on training to researchers, ensuring they can effectively analyze and interpret their metagenomic data. This personalized approach enhances researchers' proficiency in bioinformatics analysis and empowers them to extract meaningful insights from their metagenomic datasets.
Metagenomics Data Analysis Services Features
100% Post Service Assistance
Quality Assurance
Service Customization
Data Security
Experienced Team
Analysis Cases:
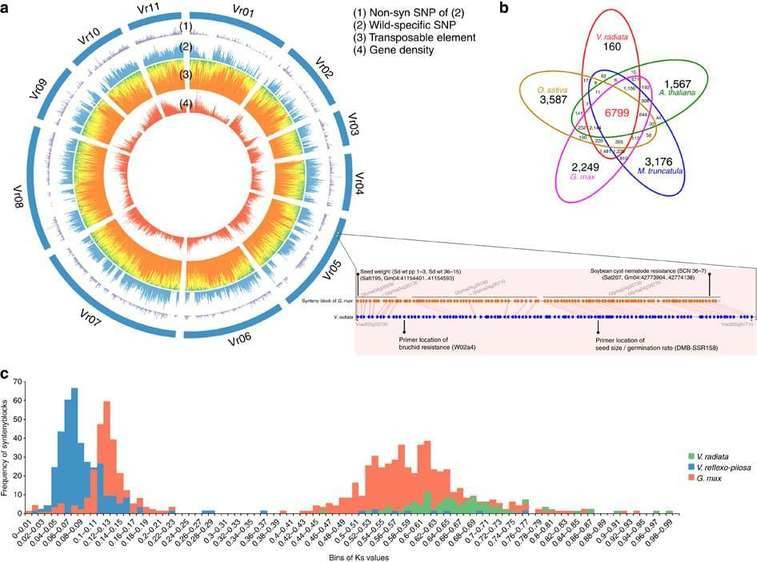
|
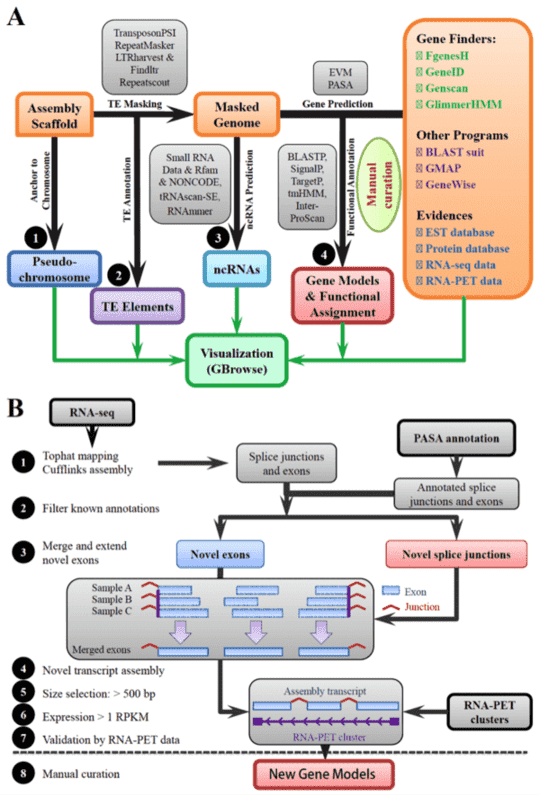
|
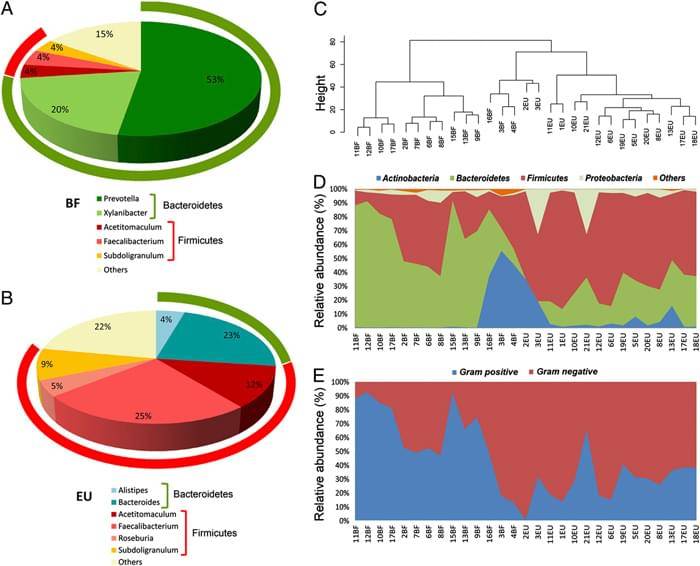
|


 Sample Submission Guidelines
Sample Submission Guidelines
