Comprehensive Profiling of Circular RNAs with Nanopore Sequencing and CIRI-Long
Jinyang Zhang, Lingling Hou, Zhenqiang Zuo, Peifeng Ji, Xiaorong Zhang, Yuanchao Xue & Fangqing Zhao
Published: March 11, 2021
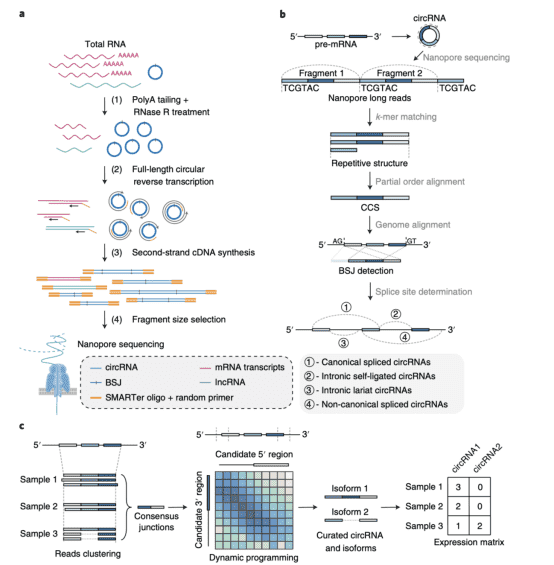
Abstract: Reconstructing the sequence of circular RNAs (circRNAs) from short RNA sequencing reads has proved challenging given the similarity of circRNAs and their corresponding linear messenger RNAs. Previous sequencing methods were unable to achieve high-throughput detection of full-length circRNAs. Here we describe a protocol for enrichment and full-length sequencing of circRNA isoforms using nanopore technology. Circular reverse transcription and size selection achieves a 20-fold higher enrichment of circRNAs from total RNA compared to previous methods. We developed an algorithm, called circRNA identifier using long-read sequencing data (CIRI-long), to reconstruct the sequence of circRNAs. The workflow was validated with simulated data and by comparison to Illumina sequencing as well as quantitative real-time RT–PCR. We used CIRI-long to analyze adult mouse brain samples and systematically profile circRNAs, including mitochondria-derived and transcriptional read-through circRNAs. We identified a new type of intronic self-ligated circRNA that exhibits special splicing and expression patterns. Our method takes advantage of nanopore long reads and enables unbiased reconstruction of full-length circRNA sequences.
More info at: https://www.nature.com/articles/s41587-021-00842-6
CIRI-long code path: https://github.com/bioinfo-biols/CIRI-long


 Sample Submission Guidelines
Sample Submission Guidelines
