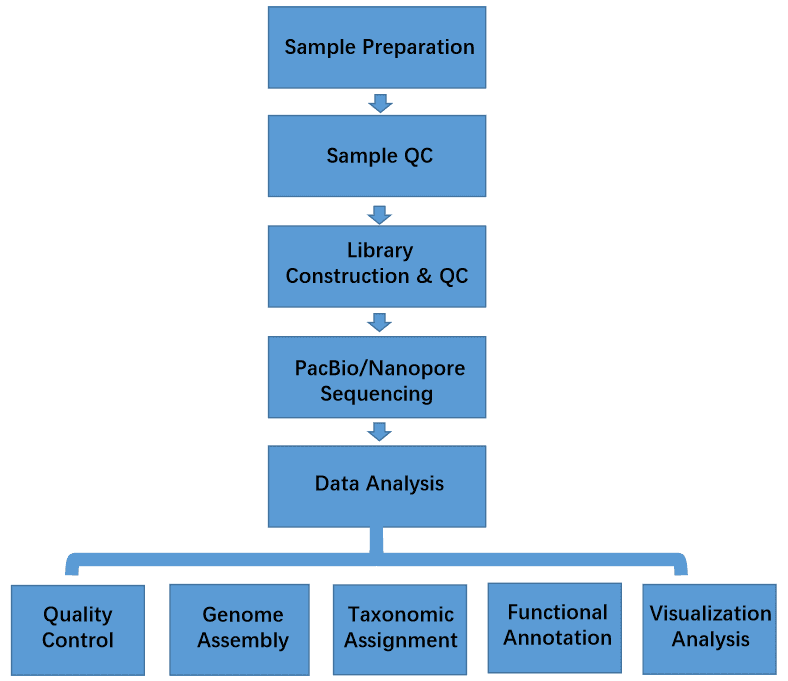
We provide the long-read metagenomics sequencing service based on the single-molecule sequencing technologies. Compared with metagenomic shotgun sequencing, the assembly of metagenomes is dramatically improved by long-read sequencing technologies including PacBio SMRT and nanopore sequencing. Chemical modifications can be directly detected without sample preprocessing for next-generation sequencing (NGS) such as bisulfite treatment of antibody pull-down. Results of metagenomic samples achieved >99% predicted concordance to reference sequences in soil, water, etc. Long-read metagenomic sequencing can detect very low-abundance members of the microbial communities that may be missed or are too expensive to identify by other approaches. They can identify rare species (low as 0.05%) to comprehensively describe the composition of a microbial community and the abundance of its members.

Coupled with powerful computational algorithms and tools, long-read metagenomics technologies could provide much important information about species-level classification, microbial community structure, the abundance of microorganisms, genetic variations, gene annotation, functional annotation, pathway enrichment, microbial evolution and so on. Metagenomics enables the detection of bacteria, archaea, viruses, and eukaryotes in samples. The long-read metagenomic sequencing can reduce assembly errors, and largely improve the resolution. Long-read metagenomics sequencing is an ideal platform for sequencing of microbial communities, particularly for the complex samples and low-diversity microbial communities.