ATAC-Seq – A Method to Study Open Chromatin
A Research into Chromatin
In eukaryote cells, the chromatin DNA replication and gene transcription occur when the higher-order structure of DNA becomes loose. Such a structure is called open chromatin, which has sufficient space to allow for binding with regulatory proteins (such as transcription factors and cofactors). Transcription regulation can be better understood at the DNA level by studying open chromatin regions in specific cell states.
What Is ATAC-Seq?
Assay for transposase-accessible chromatin with high-throughput sequencing, short for ATAC-Seq, is a relatively new technology that was introduced in 2013. ATAC-Seq utilizes hyperactive mutant Tn5 transposase that inserts sequencing adapters to accessible DNA regions, by which open chromatin sequences can be distinguished from the whole chromatin. Then through high-throughput sequencing and bioinformatics analysis, genetic information and other relevant biological issues can be analyzed and explored.
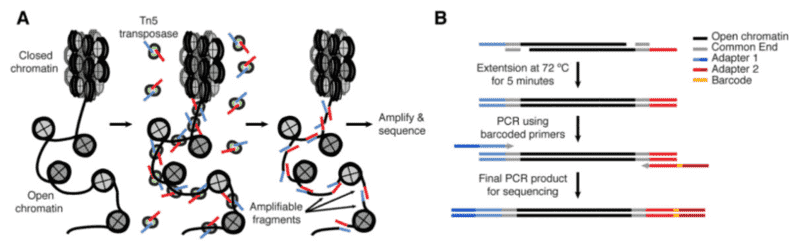
Figure 1. The basic principle of ATAC-Seq (Buenrostro 2015)
Compared to traditional methods such as MNase-Seq and DNase-Seq, ATAC-Seq has significant advantages including easier operation and higher reproducibility. And the sequencing result can be highly consistent with MNase-Seq, DNase-Seq, and Chip-Seq. Meanwhile, ATAC-Seq only requires a small amount of cells/tissue and produces better signals. It has become a preferred method to study epigenomics because it can reveal the genomic location of the open chromatin, the interaction of DNA-binding proteins, and transcriptional binding sites.
The Applications of ATAC-Seq
Eukaryotic cells assemble chromatin by folding genomic DNA and histones at multiple orders. This process plays a decisive role in gene transcriptional regulation. Chromatin accessibility is the prerequisite for the interaction between specific trans-acting factors and cis-regulating elements. Abnormal gene expression or alterations in chromatin regulatory factors will have a profound impact on cell physiology, leading to the occurrence and development of various diseases.
ATAC-Seq has now been widely used in the identification of transcriptional regulatory sequences. And it can be used in the reprogramming study for all kinds of eukaryotic cells. In the field of medicine, it can promote the research on the pathogenesis of major diseases, drug development, mechanism of action (MOA), and biomarker function. ATAC-Seq can be a versatile tool in various studies, including but not limited to:
- Mapping of chromatin accessibility sites.
- Epigenetic modification research on plants, embryo development, tumorigenesis, etc.
- Transcriptional activity research related to different tissues, organs, diseases, etc.
- Prediction of potential disease biomarkers.
- The functions of transcription factors in gene expression regulation.
- Interpretation of transcriptional regulation mechanism and genome-wide regulatory network coupled with RNA-Seq.
Despite the above advantages and wide applications that make ATAC-Seq a powerful tool in epigenomic studies, there are also other matters that one should not neglect. ATAC-Seq can be used to measure the genome-wide open chromatin to obtain possible protein binding sites, which is generally used for the sequencing of unknown transcription factors or screening for specific regulators of interest coupled with other technologies. In addition, the activity of Tn5 transposase is affected by the composition of the reaction solution and reaction conditions, which need to be optimized for chromatin shearing.
Reference:
- Buenrostro JD; et al. ATAC-seq: A Method for Assaying Chromatin Accessibility Genome-Wide. Current Protocols in Molecular Biology. 2015, 109: 21.9.1-.9.9.(1).


 Sample Submission Guidelines
Sample Submission Guidelines
