CRISPR Sequencing: Introduction, Workflow, and Applications
The CRISPR-Cas9 system
The CRISPR-Cas9 gene targeting system, which consists of two components: a custom guide RNA (sgRNA, which contains target-specific crRNA sequence and tracrRNA) and a non-specific CRISPR-associated endonuclease (Cas9), is a new genome engineering tool that allows scientists to modify parts of the genome by removing, adding, or changing a section of the DNA sequence in an organism. It is presently the most simplified, flexible, accurate, and efficient method of genetic manipulation, with numerous applications in medicine and crop seed enhancement. CRISPR-Cas9 has also revolutionized the generation of targeted mutations by allowing for high-throughput genome editing.
Introduction to CRISPR Sequencing
In the CRISPR experimentation process, validating edits is particularly crucial. Because off-target mutations, rather than edits in the target gene, can occur, next-generation sequencing (NGS) is a simple and high-throughput technique to verify for preferred mutations. For academia, clinics, and industry, CRISPR amplicon sequencing has become a standardized validation method. The idea of high-throughput CRISPR screening is focused on targeted amplicon sequencing, which is done using PCR experiments with primers flanking the focused area. The most delicate technique for checking mutations is targeted amplicon sequencing, which has detection frequencies as low as 0.01%.
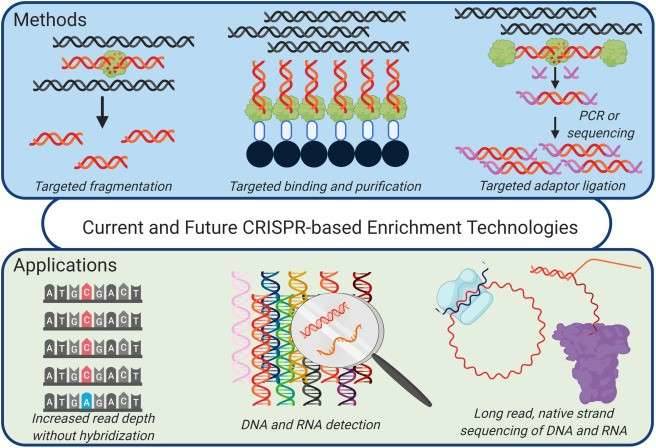 Figure 1. CRISPR-based enrichment strategies for targeted sequencing. (Schultzhaus, 2020)
Figure 1. CRISPR-based enrichment strategies for targeted sequencing. (Schultzhaus, 2020)
CRISPR sequencing can provide direct and comprehensive data about the nature and variety of the mutations, such as confirmation of knockout/knockin alleles, sgRNA cutting efficiency assessment, homozygous and heterozygous identification, mutation frequencies calculation, and so on.
CRISPR Sequencing Workflow
DNA isolation, PCR experiments, deep sequencing, and data analysis are all part of the CRISPR mutation sequencing workflow. Researchers can use CRISPR mutation sequencing to verify guides library, evaluate CRISPR/Cas9 targets and mutation efficiency, and find the most promising target sites.
Step 1: Design- Select a Cas enzyme and create a guide RNA.
Step 2: Deliver- It is recommended to combine Cas enzyme and guide RNA to form a ribonucleoprotein to deliver Cas enzyme and guide RNA to cells (RNP). Electroporation or lipofection can then be used to deliver the RNP to cells. To improve transfection efficiency, separate electroporation enhancers can be utilized for Cas9 and Cas12a.
Step 3: Repair- Cas9 or Cas12a-containing RNPs cut the genomic DNA, causing a double-strand break (DSB). Non-homologous end joining (NHEJ) and homology-directed repair (HDR) are natural processes in cells. The NHEJ pathway can be used to create knockouts in order to study gene function. HDR necessitates not only RNP to create the DSB, but also a blueprint to direct the repair, and can be employed to introduce specific mutations.
Step 4: Analyze- Depending on your research needs, you can use a variety of methods to detect mutations. Gel-based methodologies can be used to determine the presence of a transition in a genomic sequence in a non-specific manner. Next-generation sequencing (NGS) is strongly advisable for establishing the success of on-target editing and investigating off-target effects.
Applications
CRISPR-Cas9 technology has transformed the ability to modify DNA and modulate the expression levels of genes of interest, supplying effective tools to speed up the accurate engineering of a variety of organisms. Furthermore, the CRISPR-Cas system can be used to create "precision" antimicrobials that target bacterial pathogens based on their DNA sequence. By selectively targeting genes associated with antibiotic resistance, biofilm formation, and virulence, this potential will allow drug-resistant microbes to be killed.
References:
- Schultzhaus Z, Wang Z, Stenger D. CRISPR-based enrichment strategies for targeted sequencing. Biotechnology Advances. 2020 Nov 28:107672.
- de la Fuente-Núñez C, Lu TK. CRISPR-Cas9 technology: applications in genome engineering, development of sequence-specific antimicrobials, and future prospects. Integrative Biology. 2017 Feb 1;9(2).
- Selle K, Barrangou R. CRISPR‐Based technologies and the future of food science. Journal of food science. 2015 Nov;80(11).
- Makarova KS, Wolf YI, Alkhnbashi OS, et al. An updated evolutionary classification of CRISPR–Cas systems. Nature Reviews Microbiology. 2015 Nov;13(11).


 Sample Submission Guidelines
Sample Submission Guidelines
