Current Techniques for Targeted Region Enrichment
What is targeted region sequencing?
While whole genome sequencing covers the entire genome of an organism, targeted region sequencing determines the order of nucleotides in the regions of interest. Utilizing targeted genome sequencing, you can detect SNPs, indels, CNVs, and large SVs in a highly sensitive and specific manner. Compared to whole genome sequencing, targeted genome sequencing is more focused, economical, and effective. Furthermore, targeted regions can be sequenced at a much greater depth for a lower cost, which contributes to detecting rare variations. Whole exome sequencing is an example of targeted genome sequencing since it focuses on the exons. And using a targeted or “hot-spot” panel, we can sequence an array of genes of interest. These genes may harbor mutations that are associated with the pathogenesis of diseases, or these genes are clinically-actionable genes of interest. This has been applied to biological and medical research, as well as clinical care. Here we mainly discuss the current target enrichment techniques and recommend qualified products.
Classical techniques for targeted enrichment
Target enrichment greatly facilitates the development of targeted region sequencing, enabling sequencing affordable and efficient for complex genomes. There are three classical strategies for targeted enrichment (Figure 1).
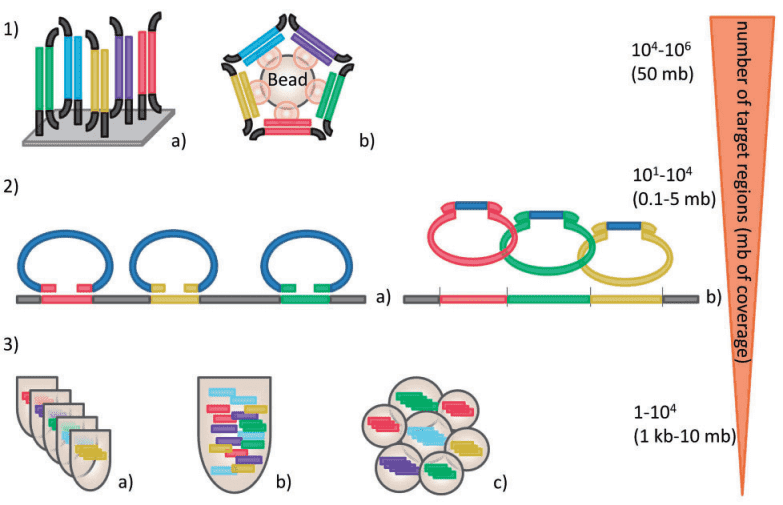
Figure 1. Strategies for targeted enrichment (Mertes et al. 2011). 1) Hybrid capture either on microarrays (a) or in solution (b). 2) Enrichment by MIPs (a), or selector probes (b). 3) Enrichment by typical PCR (a), multiplex PCR assay (b), and RainDance micro droplet PCR with up to 20,000 primer pairs (c).
- Hybrid capture
The shotgun fragment library is prepared to hybridize to a library that contains DNA fragments complementary to the target regions. This can be performed on microarrays or in solutions. Mamanova et al. (2010) found that for small target sizes (approximately 3.5 Mb), solution capture yields superior coverage of the target regions than array-based approach, and that for whole exome enrichment, both approaches perform equivalently. We recommend SureSlect (Agilent Technologies), Nextera (Illumina), TruSeq (Illumina), and SeqCap EZ (Roche NimbleGen) for in-solution hybridization-based target enrichment.
- Selective circularization
Molecular inversion probes (MIPs) consist of a common sequence flanked by target specific sequences. After the hybridization to the regions of interest, MIP is subjected to a gap filling reaction and ligation to generate closed circles. The MIPs can be hybridized to mechanically sheared DNA, while selector probes use a restriction enzyme cocktail to digest the DNA and the probes are adapted to the restriction pattern. MIPs and selectors consist of a common central linker, the sequencing primer for NGS can be included into this linker. Therefore, NGS library construction is not required. We recommend HaloPlex (Agilent Technologies) for selective circularization-based target enrichment.
- PCR amplification
PCR is used to enrich regions of interest by conducting multiple long-range PCRs in parallel, multiplex PCRs, and microdroplet PCRs. There are many available products for long-range PCR, including SequalPrep (Thermo Fisher Scientific) and SeqTarget (Qiagen), Ion Ampliseq based on high-multiplex PCR, and RainDance’s microdroplet technology.
The comparisons of classical target enrichment techniques
The four strategies have its own merits and demerits. For example, the specificity of PCR exceeds that of hybrid capture, but its uniformity is not matched by either MIPs or hybrid capture. For updates of protocols, you can visit this website: ftp://ftp.sanger.ac.uk/pub/pulldown/.
Table 1. Comparisons of target-enrichment methods. Adapted from Dapprich et al. (2016).
| Advantages | Limitations | Coverage uniformity of target bases | On target reads (%) | Cost per amplicon | |
| Traditional/ Multiplex PCR |
|
|
80-100% at 100X | 95 | $5.48 (Sanger sequencing) |
| Microdroplet PCR |
|
|
98.45-100% | 52.5 | $1.56 |
| Hybridization capture |
|
|
92.96-100% | 53.3-60.7 | $2.34 |
| Selective circularization |
|
|
- | - | - |
Advanced method: Region-Specific Extraction (RSE)
The traditional methods heavily rely on fragmentation of DNA prior to amplification, which generate relatively short sequences (less than 1kb). This is a limiting factor for characterizing complex genomic loci since they could not provide fragments large enough to span confounding sequence elements. Fortunately, the enrichment method — Region Specific Extraction (RSE), proposed by Dapprich et al. (2016), addresses this unmet need by capturing long DNA fragments (approximately 20kb).
The principle of RSE is outlined in Figure 2. Briefly, the genomic DNA is denatured to be hybridized with capture primers. The bound primers are then enzymatically extended with incorporated biotinylated-dNTPs. The targeted DNA segments are pulled down by streptavidin-coated magnetic particles. The captured DNA can be amplified by whole genome amplification and subsequently processed by next-generation sequencing.
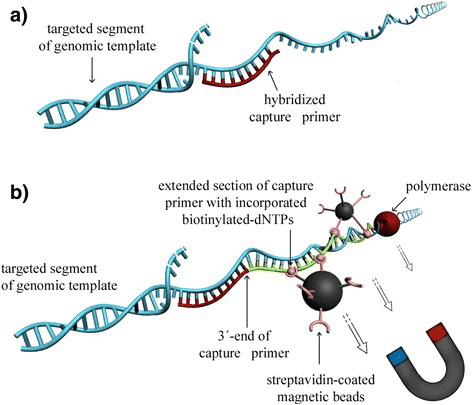
Figure 2. Principle of RSE.
Metrics for the assessment of a target-enrichment experiment
There are several metrics need to be considered in target-enrichment experiments. To deal with this issue, a universal set of Target Enrichment Sequencing Descriptors (TESD) metrics has been developed, enabling to assess target enrichment results. The TESD uses the following parameters:
- Region of interest, ROI
- Average read depth in region of read, DROI
- Specificity, S
- Enrichment factor, EF
- Evenness, E
- Weight of input DNA requirement, W
- Fraction sufficiently covered at a read depth of x, Fx
Of the seven metrics, ROI and W determine the input requirement, and the other five are a measurable report on either the enrichment (S, EF, E) or on the sequencing results (DROI, Fx).
If you are interested in our genomics services, please visit our website: www.cd-genomics.com for more information. We can provide a full package of genomics sequencing, including whole genome sequencing, whole exome sequencing, targeted region sequencing, mitochondrial DNA (mtDNA) sequencing, and complete plasmid DNA sequencing.
References:
- Dapprich J, Ferriola D, Mackiewicz K, et al. The next generation of target capture technologies-large DNA fragment enrichment and sequencing determines regional genomic variation of high complexity. BMC genomics, 2016, 17(1): 486.
- Mamanova L, Coffey A J, Scott C E, et al. Target-enrichment strategies for next-generation sequencing. Nature methods, 2010, 7(2): 111.
- Mertes F, ElSharawy A, Sauer S, et al. Targeted enrichment of genomic DNA regions for next-generation sequencing. Briefings in functional genomics, 2011, 10(6): 374-386.
- McGinn S, Bauer D, Brefort T, et al. New technologies for DNA analysis–a review of the READNA Project. New biotechnology, 2016, 33(3): 311-330.
- Ballester, L. Y., Luthra, R., Kanagal-Shamanna, R., et al. Advances in clinical next-generation sequencing: target enrichment and sequencing technologies. Expert Review of Molecular Diagnostics, 2016, 16(3), 357–372. doi:10.1586/14737159.2016.1133298


 Sample Submission Guidelines
Sample Submission Guidelines
