Methylation Panel: Unlocking the Research Potential of Epigenetics
DNA methylation serves as a critical epigenetic regulatory mechanism in humans, functioning like genetic "switches" that control gene activity. Our Methylation Panel series comprises precision testing toolkits utilizing high-throughput sequencing (NGS) or advanced molecular detection technologies. By precisely locating and analyzing methylation changes in targeted genomic regions (e.g., promoter regions, CpG islands), these panels reveal epigenetic signatures closely linked to disease pathogenesis (particularly oncology), aging processes, and individual responses to pharmaceuticals/environmental factors. Essentially, they decode pivotal regulatory signals impacting health and disease – independent of genetic sequence alterations.
Key Features:
- Comprehensive Coverage, Optimized Targets
Featuring modular panel designs, the series addresses diverse high-value research scenarios. All targets undergo rigorous selection and scientific validation, leveraging large-scale datasets and cutting-edge research evidence to ensure scientific validity.
- Advanced Technology, Precision Reliability
Employing proprietary methylation capture technology, our panels deliver high sensitivity, specificity, and reproducibility – detecting subtle methylation changes at single-base resolution.
Request a Quote
What is targeted methyl-seq?
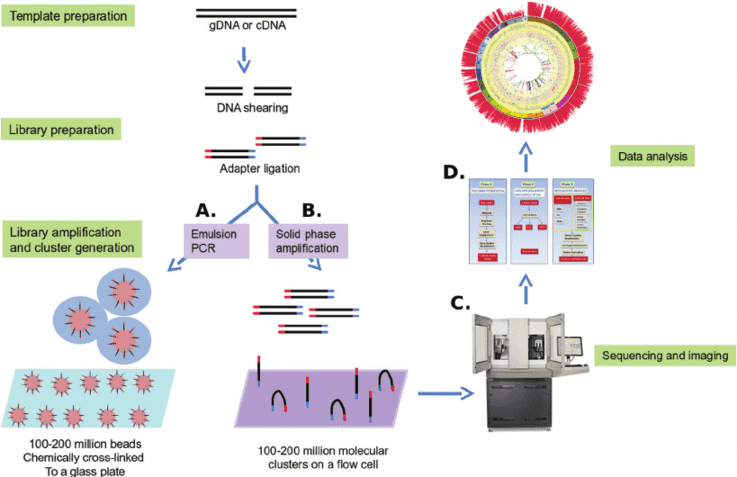
Targeted Methylation Sequencing (Targeted Methyl-Seq), also known as Targeted Bisulfite Sequencing, is a next-generation sequencing (NGS) methodology designed to precisely analyze DNA methylation patterns. This epigenetic modification provides critical insights for research areas like cell type identification in epigenetics and oncology. While whole-genome bisulfite sequencing (WGBS) assesses methylation across the entire genome, Targeted Methyl-Seq utilizes hybridization probes to selectively capture and enrich specific, biologically relevant genomic regions. This focused approach enables significantly deeper sequencing depth and higher coverage of the targeted areas compared to WGBS, offering enhanced resolution for regions of particular interest.
How Does Bisulfite Sequencing Work?
Bisulfite sequencing leverages the power and sensitivity of NGS for methylation analysis. The core principle relies on the chemical conversion of DNA using bisulfite treatment. During library preparation, this process specifically converts unmethylated cytosines (C) to uracil (U). Subsequent PCR amplification and sequencing then identify these converted bases as thymine (T) in the resulting data. By quantifying the proportion of cytosines remaining as C (indicating methylation) versus those read as T (indicating original unmethylation) at specific genomic positions, researchers can determine the precise percentage of methylated cytosines.
Bisulfite sequencing can be applied comprehensively via WGBS or focused on defined regions using targeted methods like Target Enrichment Methyl-Seq or Amplicon Methyl-Seq. Furthermore, specialized techniques such as Oxidative Bisulfite Sequencing (OxBS) and TET-Assisted Bisulfite Sequencing (TAB-Seq) can be integrated with NGS. These complementary chemistries allow for the specific identification and differentiation of hydroxymethylcytosine (5-hmC) alongside the analysis of standard methylcytidine (5-mC).
Proprietary Probe Design for Methyl-seq

This solution targets gene regulatory regions (CGI, 5' UTR, Promoter, Silencer, etc.), enabling dual-state detection for CpG sites on both DNA strands. It requires only twice the probe consumption of conventional panels, ensuring high methylation detection rates while significantly saving probes and reducing costs.
Workflow of Targeted Methyl-seq

Why Choose CD Genomics
Trusted Quality Control: Our multi-layered QC system, implemented across the entire workflow, guarantees every step – from raw materials to final products – meets the highest standards. This ensures reliable product performance, providing robust support for scientific discovery and health research.
One-Stop Solution: We deliver a comprehensive suite including sample processing, DNA extraction, library preparation, hybrid capture kits, and supporting professional bioinformatics analysis and interpretation.
Expert Support and Partnership: Access the expertise of our experienced epigenetics specialists and bioinformatics team. We provide dedicated technical consultation and foster ongoing collaborative research partnerships.
* For research purposes only, not intended for clinical diagnosis, treatment, or individual health assessments.
Related Services
Related Products
Related Resources
Explore Our Methylation:
| Cat. No. |
Product Name |
Brief Description |
Inquiry |
Basket |
| PNM001 |
CDCap™ Human CpG Island Panel Kit |
CDCap™ Human CpG Island Panel Kit enables precise DNA methylation detection across 27,000 CpG islands and 13 markers in human genomic DNA. |
|
|
| PNM002 |
CDCap™ Early Methylation Screening Panel Kit |
CDCap™ Early Methylation Screening Panel: RUO tool detecting DNA methylation variants via liquid biopsy. Targets 12 biomarkers across 8 cancers using sensitive library prep for non-invasive early detection research. |
|
|