Hybridization Capture-Based NGS Panel
CD-Genomics is a company with rich experience in targeted gene panel sequencing which is used to accurately detect gene variations related to diseases.
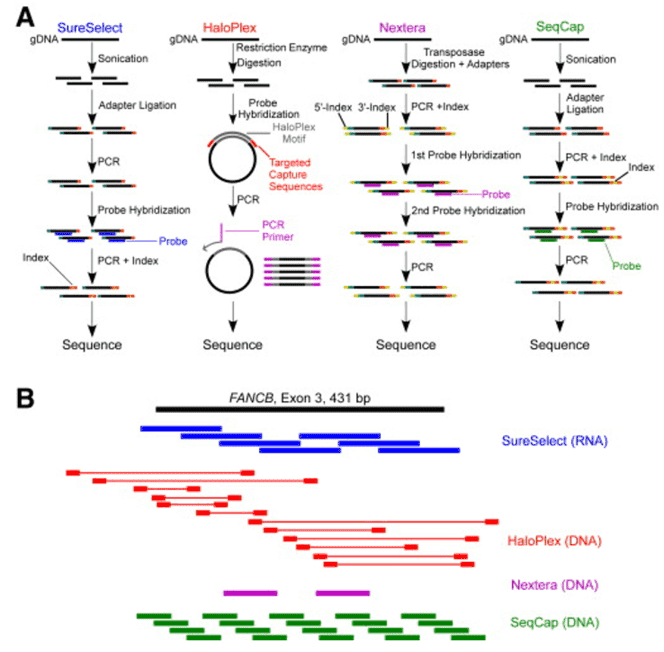
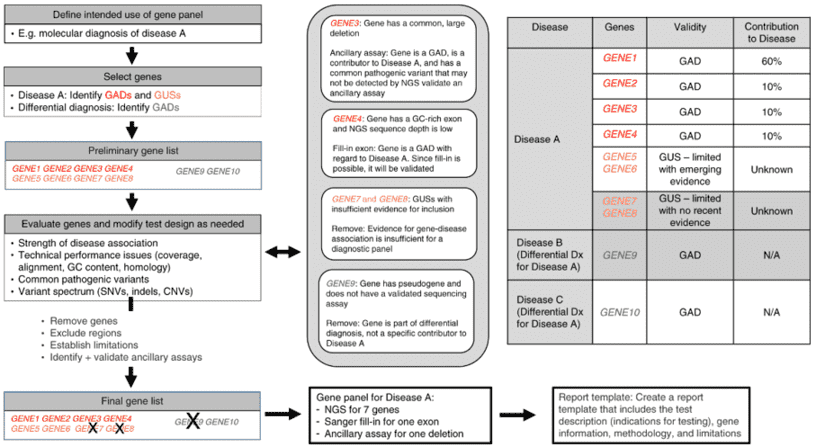
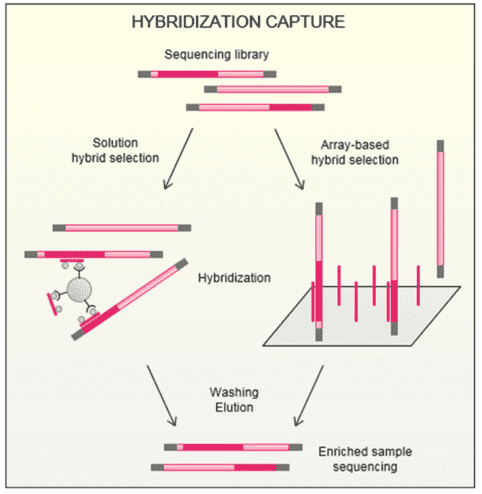
CD-Genomics provides custom NGS panels for mutations in at least 68 diseases and thousands of genes. We can employ two target enrichment strategies prior to the NGS panel, which are amplification and hybridization capture. In a hybridization capture strategy, the ‘capture’ can be realized through the hybridization of fragmented DNA with baits complementary to regions of interest, either on a microarray chip or free in solution. In-solution hybridization capture can be scalable and easily automatable and rules out the shortcomings of microarray hybridization such as high cost, limitations to the number of samples and large sample requirements. Thus, solution-based hybridization becomes the mainstream approach of capture-based enrichment for NGS panel.
Highlight Applications
For larger disease panels
Disease-related gene discovery
Rare disease-related variant detection
Widely used in oncology research, inherited disease research, and more
Hybridization Capture-Based Disease Panel Workflow
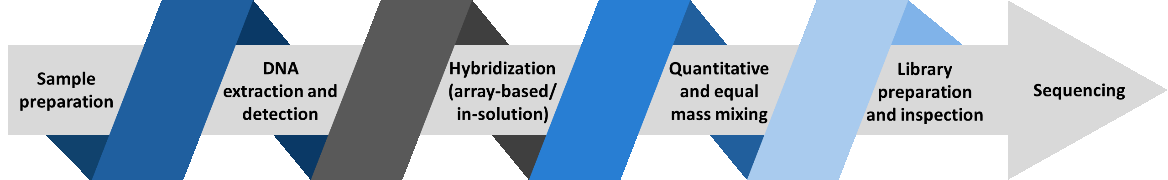
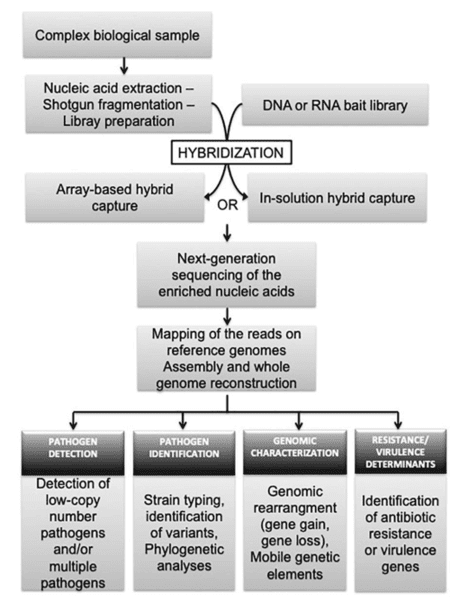
In a hybridization assay, the protocols start with random shearing of the genes, which are later denatured by heating. Randomly sheared overlapping fragments are captured by DNA or RNA single-stranded oligonucleotides specific to the region of interest. DNA bound to the baits is separated from background DNA. The baits are removed and DNA enriched for regions of interest is taken forward to the second round of PCR, followed by NGS of the enriched nucleic acids.
Our strict quality control throughout the pipeline workflow to ensure the accuracy and repeatability of the enrichment, sequencing and bioinformatics analysis of hybridization capture-based NGS panel.
Features of Our Custom Hybridization Capture NGS Panel
Highly accurate hybridization-based NGS panel for scientific research purposes
High sensitivity for variant calling
Provides depth and uniformity of coverage required for genetic variant discovery studies
Focusing on specific regions of interest – efficient and cost-effective
Sample Requirements
- DNA > 1 μg, no obvious degradation, 1.8 < OD260/280 < 2.0, OD260/230 ≥ 1.8, concentration ≥ 30 ng/μL.
-
Or properly treated blood or tissue samples.
Bioinformatics Analysis
| Analysis Contents |
Details |
| Bait design |
The design of nucleic acid baits for hybridization capture. |
| Primary analysis |
To identify single nucleotide variants and small insertions/deletions. |
| Trimming pipeline |
Quality control check is implemented and only samples with enough reads and base quality are considered |
| Variant identification & annotation |
Mutations are considered pathogenic if they are absent from controls, predicted to alter the sequence of the encoded protein and to adversely affect protein function. |
| Allelic variant profiling |
To detect and characterize complex allelic variants, such as short tandem repeats. |
| Variant classification |
To detect mutations at low-allelic fraction from sequencing of matched tumor-normal sample pairs. |
Why choose us?
- Advanced technology platform and experienced team of scientists
- Customized services for the detection of diverse disease genes
- Bioinformaticians can get you more valuable information from the NGS panel
- Affordable price and fast turnaround time
Hybridization-based assays may offer wider scope for superior performance through optimization of bait design. It works well for disease-related rare variant detection including single nucleotide polymorphism detection, indel detection, copy number variation detection, and structural variation detection. Select and submit the target genes you are interested in and we will send you a design coverage report and provide you with a quote. If you have any questions or requirements, please feel free to contact us, we will help you solve the problems and further optimize to meet your needs.
* For research purposes only, not intended for clinical diagnosis, treatment, or individual health assessments.
Related Services
Related Products
Related Resources