Overview of Gene Target Enrichment Methods for Disease Research
Development of Target Enrichment
Target Enrichment is a stage of pre-sequencing preparation of DNA where it can be directly amplified (amplicon or multiplex PCR-based) or captured (hybrid capture-based) DNA sequence. Using DNA sequencers, these enhanced DNA fragments can then be sequenced. The process for Targeted Sequencing, also known as Resequencing, is called Target Enrichment. With target enrichment, only a portion of the whole genome, or regions of interest, is amplified and sequenced without sequencing the whole genome of a specimen. Target enrichment is needed in order to concentrate only on the particular area of a genome or DNA specimen, for the research of various diseases such as pathogenic diseases, inherited diseases, and cancer.
In its initial hybridization method, the hybridization method focused on capturing DNA on a solid surface on a single microarray with 385,000 probes. While later models had more sophisticated capabilities, array-based methods were less common than solution-based capture methods developed subsequently because of the need for large quantities of DNA and expensive hardware and its reduced throughput. There are many target enrichment methods available. Sensitivity, accuracy, uniformity of coverage, and reproducibility are important criteria that may also play a role in selecting the target enrichment variable. The choice of which tool to use depends on a variety of factors such as price, ease of usage, and the amount of sample input accessible. Hybridization-based and PCR-based target enrichment approaches are some primary target enrichment strategies. DNA is broken (by physical shearing or enzymatic techniques) in the Hybridization-based process and primed for sequencing by inserting adapters unique to the sequencing platform used, usually serving as barcodes for later detection. While amplicon-based or PCR-based approaches are used to amplify particular regions of interest by primers. Tens of thousands of primer pairs are being used for PCR reactions, allowing many regions of interest to be targeted simultaneously and requiring a small amount of DNA input.
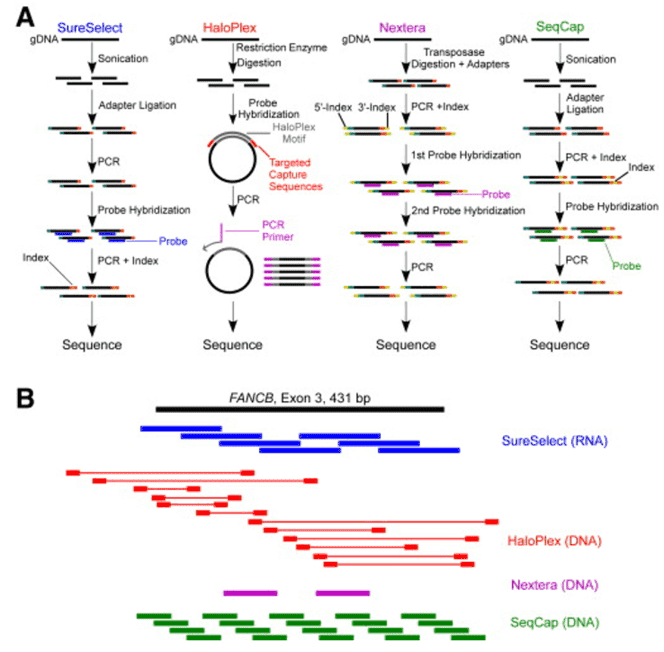 Figure 1. Methods summary and capture methods for targeted DNA sequencing. (Samorodnitsky, 2015)
Figure 1. Methods summary and capture methods for targeted DNA sequencing. (Samorodnitsky, 2015)
Advantages of Target Enrichment Method
Targeted sequencing offers the opportunity to concentrate on particular disease-associated genes or other specific genes of concern as a targeted sequencing technique. With this method, it is possible to ignore the rest of the entire genome, streamlining subsequent bioinformatics research and having the opportunity to achieve broader coverage. Coverage depth (redundancy of coverage) matters as it can increase confidence in the simple call for variant analysis. For targeted sequencing, target enrichment, an additive phase to sample preparation, is necessary and can be done by a combination of methods. Target enrichment is highly beneficial for getting the necessary read depth and consistency of coverage in genetic variant discovery studies. Using target enrichment methods, on-target reads and reads matched to a genome can be fully utilized. Time and sequencing strategies will be focused on the same disease area of research concern with the accessibility of targeted panels for human illnesses from multiple vendors. Some targeted modules may also be custom-made, making it easy to concentrate on regions of concern. In analyzing variants of unknown importance, a targeted approach is often useful as the research may be based on a small amount of data.
References:
- Kozarewa I, Armisen J, Gardner AF, et al. Overview of target enrichment strategies. Current protocols in molecular biology. 2015, 112(1):7-21.
- Samorodnitsky E, Datta J, Jewell BM, et al. Comparison of custom capture for targeted next-generation DNA sequencing. The Journal of Molecular Diagnostics. 2015, 17(1):64-75.
- Mamanova L, Coffey A, Scott CE, et al. Target enrichment strategies for next-generation sequencing. Nature Methods. 2010, 7;2:111-118.
* For research purposes only, not intended for clinical diagnosis, treatment, or individual health assessments.
Related Services
Related Products
 Figure 1. Methods summary and capture methods for targeted DNA sequencing. (Samorodnitsky, 2015)
Figure 1. Methods summary and capture methods for targeted DNA sequencing. (Samorodnitsky, 2015)