- Home
- Products
- Ready-to-Use NGS Panel
- Real-Time PCR Kits
- Discover the infinite power of our Respiratory Virus Kits to drive scientific discovery
- Cancer Real-Time PCR Kits
- Pathogens Detection
- Respiratory Pathogen RT-PCR Kits
- Respiratory Virus RT-PCR Kits
- EBV RT-PCR Kits
- HBV RT-PCR Kits
- HPV RT-PCR Kits
- HLA Typing Kits
- Food & Environmental Microbiology Test Kits
- Pharmacogenomics Testing Kits
- Allergens Detection Kits
- Food & Animal Feed Ingredients Authentication Kits
- Plant GMO Detection Kits
- Oncology Quality Control Products
- Pathogenic Microorganisms Control Products
- QC & RM for Oncology Assays
- Copy Number Variation (CNV) Reference Products
- ctDNA (Circulating Tumor DNA) Reference Products
- Tumor Companion Diagnostic Reference Products
- Homologous Recombination Deficiency (HRD) Reference Products
- Minimal Residual Disease (MRD) Reference Products
- Homologous Recombination Repair (HRR) Reference Products
- Tumor Mutation Burden (TMB) Reference Products
- Microsatellite Instability (MSI) Reference Products
- Reproductive Health Reference Products
- Infectious Diseases Reference Products
- Inherited Diseases Reference Products
- Services
- Predesigned NGS Panel
- Inherited Disease Panel
- Discovery Cancer Panels
- Pan-Cancer Panel Sequencing
- Hereditary Cancer Panel Sequencing
- Cancer Hotspot Panel Sequencing
- Lung Cancer Panel Sequencing
- Breast Cancer Panel Sequencing
- Ovarian Cancer Panel Sequencing
- Thyroid Carcinoma Panel Sequencing
- Esophageal Cancer Panel Sequencing
- Glioma Gene Panel Sequencing
- Colorectal Cancer Panel Sequencing
- Pharmacogenomics Testing
- Pathogens Detection
- Tumor Mutational Burden Analysis
- Solid Tumor Sequencing Service
- Non-NGS Panel
- Custom NGS Panel
- Bioinformatics Analysis Service
- Gene Fusion Detection by Sequencing
- Microsatellite Instability (MSI) Analysis
- Rare Disease Genomics
- SNP Panel & Sequencing
- Predesigned NGS Panel
- Resource
- Company
- Online Inquiry
- Order Online
- Get a Quote
x

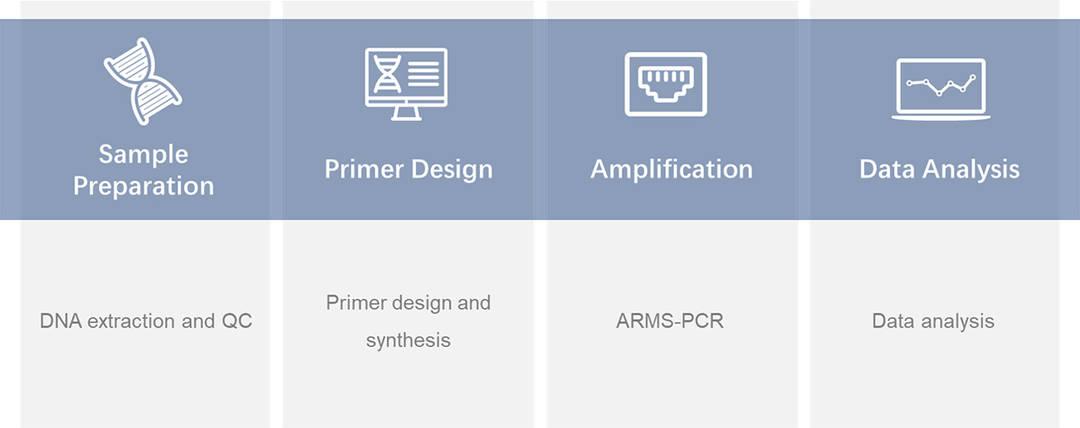 CD Genomics ARMS-PCR Service
CD Genomics ARMS-PCR Service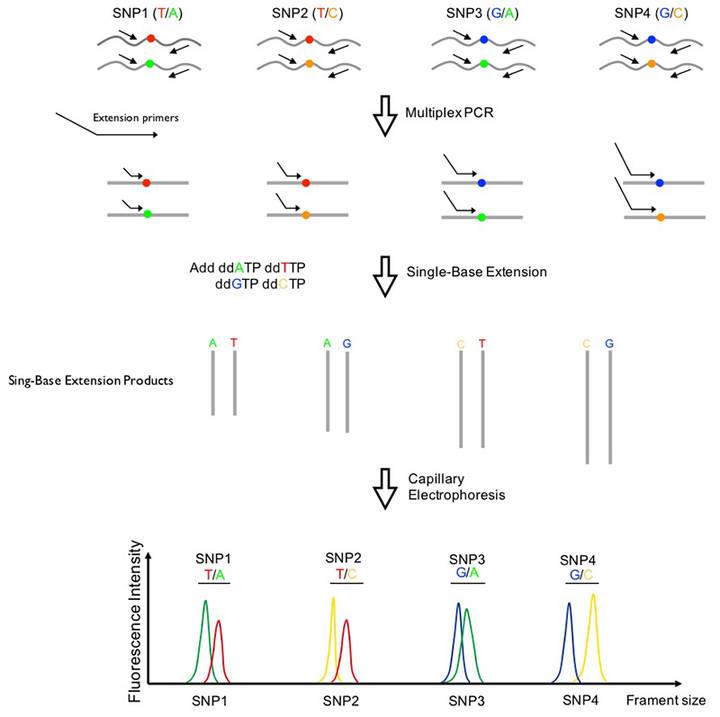 The principle and process of SNaPshot Multiplex System.
The principle and process of SNaPshot Multiplex System.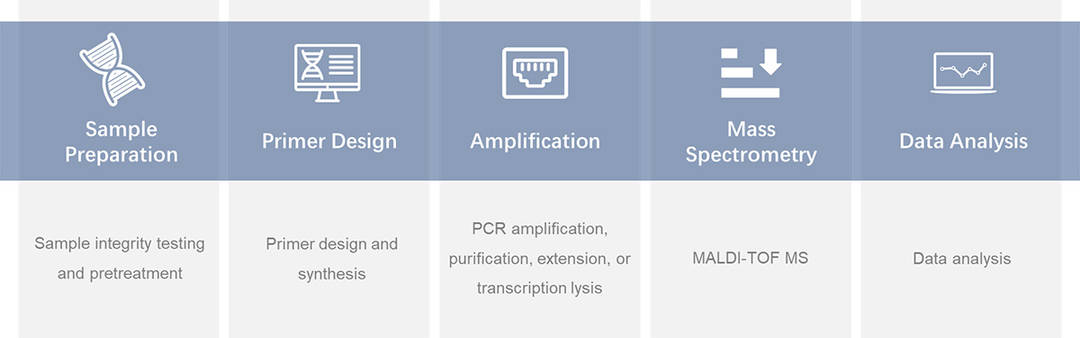 CD Genomics MassARRAY service
CD Genomics MassARRAY service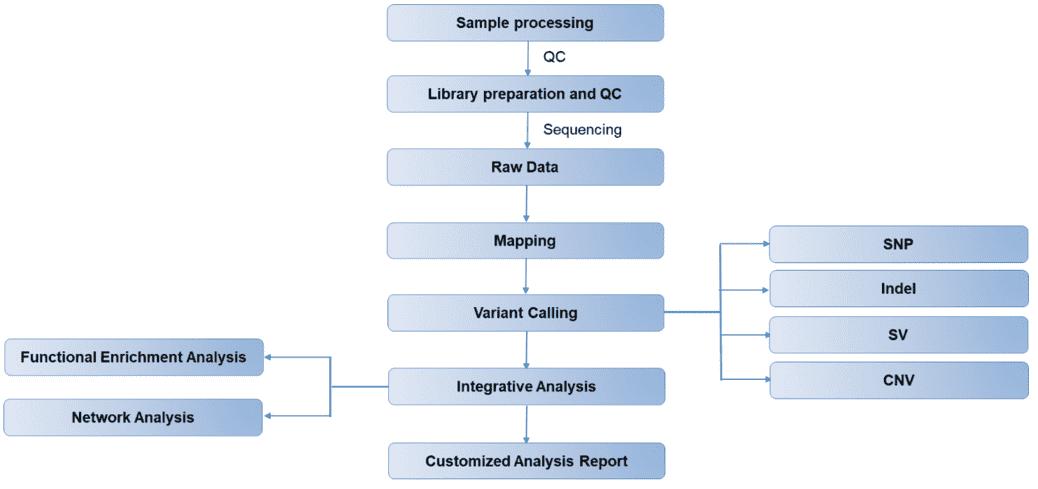 CD Genomics Solid Tumor Whole Genome Sequencing
CD Genomics Solid Tumor Whole Genome Sequencing