Custom Retinitis Pigmentosa Panel
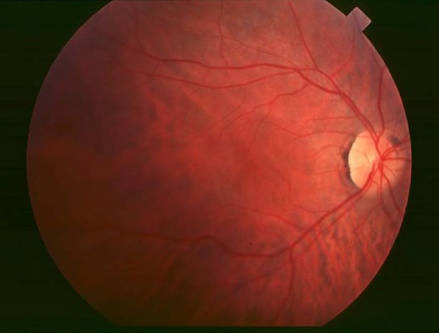
Introduction
Retinitis pigmentosa (RP) is described as a kind of retinal degeneration disease which often causes progressive loss of photoreceptors. This disease represents a retinal dystrophy with an extremely complex pathogenesis, and the symptoms are further worsened with the impairment of the retinal vascular supply. The prevalence of retinitis pigmentosa is approximately about 1:4000. It is often associated with the night blindness and eventually blindness after several decades. Up to date, there are numerous genetic mutations been identified as the causes of RP, which can explain the heterogeneity of the disease. MERTK gene mutation is a common cause of RP.
Disease-related gene description
MERTK, located on chr2q13, is responsible for the efficient phagocytosis of shed photoreceptor outer segments (POS) by the retinal pigment epithelium. Mutations in the ERTK gene disrupt phagocytosis, leading to the accumulation of exfoliative POS and the formation of subretinal debris, which results in the formation of RP. The mutations of MERTK gene account for approximately 1% to 2.5% in all the RP cases. PRPF31 is mapped on chr19q13.42. The mutations in PRPF31 can cause retinitis pigmentosa by haploinsufficiency. PRPF31 is found to account for 2.5% of autosomal dominant retinitis pigmentosa. Most PRPF31 pathogenic mutations are single-base changes or deletions that leads to premature termination of codons and nonsense mediated mRNA decay. Otherwise, PRPF31, PDE6 (PDE6A, PDE6B, PDE6G), RP25 and RPE65 also have higher prevalence in RP cases.
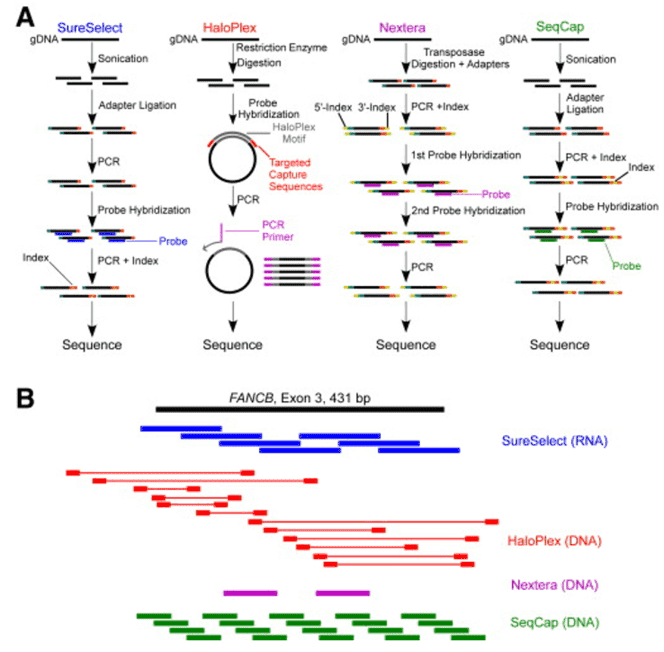
To better understand the consequences of these mutations of MERTK and other associated genes, our platform provides targeted DNA sequencing by the Illumina MiSeq or Ion PGM system, and offers a comprehensive retinitis pigmentosa panel library from which you can choose for genetic testing of retinitis pigmentosa.
Custom retinitis pigmentosa panel offers but are not limited to:
- Target enrichment with the Illumina MiSeq system is provided to efficiently and quickly detect target mutations.
- Low input DNA requirement.
- Careful construction of effective and scientifically justified gene panels is provided.
- The workflow can increase throughput and only sequence the customizable retinitis pigmentosa panel under your requirements.
- The details of test performance will be available.
- Our platform provides individual designing on custom panel or you can choose a panel from our retinitis pigmentosa library.
- The design of our custom platform is based on the frontiers from current literature about retinitis pigmentosa to target all relevant regions.
Choose the genes that suit you from the retinitis pigmentosa gene list
| ABCA4 |
ABHD12 |
AGBL5 |
AIPL1 |
ARHGEF18 |
ARL6 |
| BBS2 |
BEST1 |
C2ORF71 |
C8ORF37 |
CA4 |
RCDHR1 |
| CERKL |
CLRN1 |
CNGA1 |
CNGB1 |
CRB1 |
CWC27 |
| CYP4V2 |
DHDDS |
DHX38 |
FAM161A |
FLVCR1 |
IDH3B |
| IMPG2 |
KIZ |
KLHL7 |
LRAT |
MAK |
MERTK |
| MFRP |
NR2E3 |
NRL |
PDE6A |
PDE6B |
PDE6G |
| PEX7 |
PHYH |
PRCD |
PRPF |
RBP4 |
RDH12 |
| RDH5 |
REEP6 |
RGR |
RHO |
ROM |
RP1 |
| RP2 |
RPE65 |
RPGR |
RS1 |
SAG |
SAMD11 |
| SEMA4A |
SLC7A14 |
SNRNP200 |
SPATA7 |
SPP2 |
TTC8 |
| TULP1 |
TOPORS |
USH2A |
WDR19 |
ZNF408 |
ZNF513 |
Specimen requirements of our custom retinitis pigmentosa panel
- Specimen: whole blood, extracted DNA (no FFPE) and saliva.
- Volume: 1ng DNA or 1ng RNA.
- Collection: blood is collected by routine blood collection and saliva is collected by spitting into the provided container. DNA samples are stored in TE buffer or equivalent.
- Container: lavender-top (EDTA) tube or yellow-top (ACD) tube.
Gene panel workflow
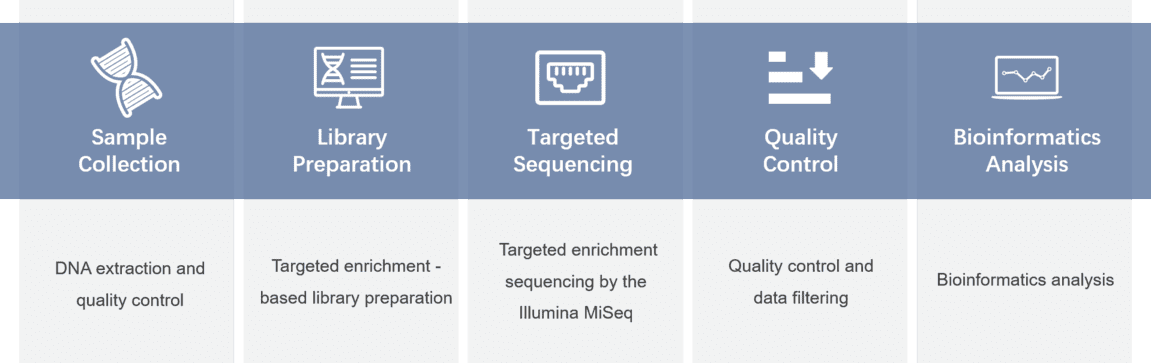
For more information about the Custom Retinitis Pigmentosa Panel or need other amplification requirements, please contact us.
References:
- Tsang SH, Sharma T. Retinitis Pigmentosa (Non-syndromic). Adv Exp Med Biol. 2018; 1085:125-130.
- He Y, Zhang Y, et al. Recent advances in treatment of retinitis pigmentosa. Curr Stem Cell Res Ther. 2015;10(3):258-65.
- Hafler BP, Comander J, et al. Course of Ocular Function in PRPF31 Retinitis Pigmentosa. Semin Ophthalmol. 2016;31(1-2):49-52.
* For research purposes only, not intended for clinical diagnosis, treatment, or individual health assessments.
Related Services
Related Products
Related Resources