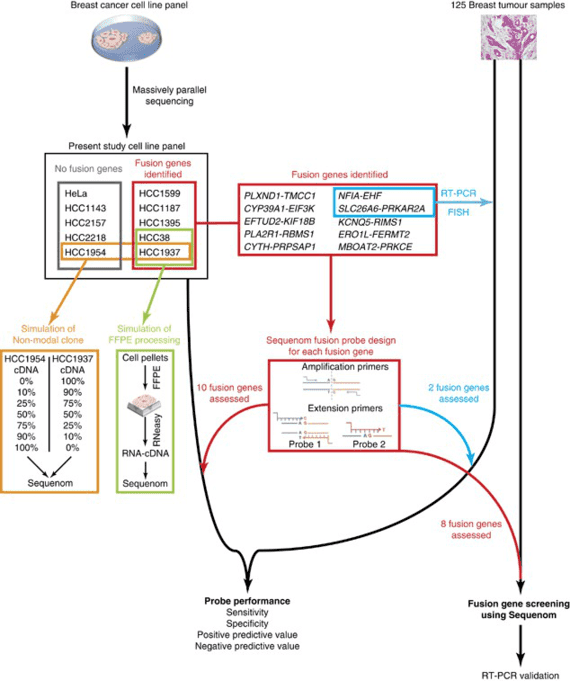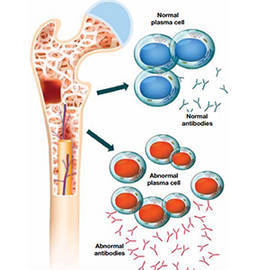
Introduction
Myeloma, also known as multiple myeloma, is divided into various kinds of types including acute myeloid leukemia (AML), myeloid dysplastic syndrome (MDS), myeloproliferative neoplasms (MPN), chronic myeloid leukemia (CML), chronic myelomonocytic leukemia (CMML) and juvenile myelomonocytic leukemia (JMML). Myeloma is a clonal plasma cell malignancy and characterized by clonal hyperplasia of malignant plasma cells in bone marrow, which accounts for 1% of all cancers and approximately 10% of all hematologic malignancies. Notably, Myeloma usually occurs in the elderly population aged 50-70 and as the population ages, its incidence is gradually increasing. Sometimes myeloma does not cause any symptoms. When myeloma is more severe, symptoms of myeloma may include bone pain, especially in the back or ribs; soft bones; fever with no certain reason; frequent infections; bruising or bleeding easily; trouble breathing; weakness of the arms or legs; and tiredness.
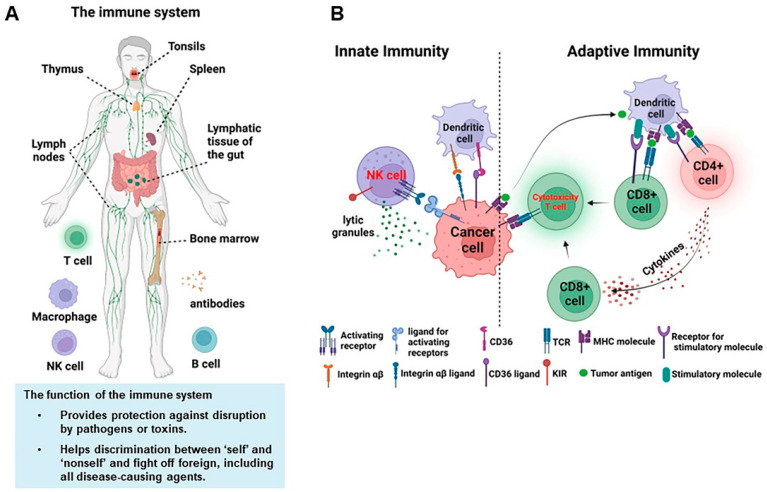
Disease-related gene description
ASXL1 is an enhancer of the trithorax and polycomb families and plays an important role in maintaining gene expression homeostasis. In recent years, ASXL1 mutations have been detected in hematopoietic cells of a variety of myeloid tumors, meaning that it plays an important role in the pathogenesis and malignant transformation of myeloid. In addition, many factors involved in regulating life activities are also associated with the development of different types of myelomas. For example, scientists have discovered that many genes, such as DNA methyltransferase 3A (DNMT3A), TET2 and EZH2 that regulate DNA methylation can contribute to AML. What's more, genes associated with RNA splicing mechanisms, including SF3B1, U2AF1, ZRSR2, SRSF2, PRPF40B, SF3A1, etc. increase the risk of myeloma. And CXCR4, CTLA-4, PD-1, PD-L1, etc. are all research hotspots.
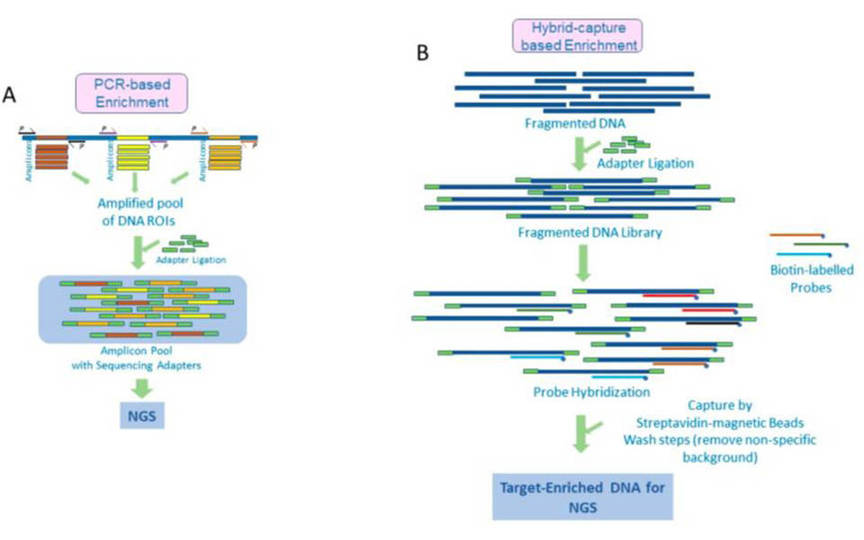
To meet clinical researches related to myeloma and other related genes, our custom myeloid panel platform provides targeted DNA sequencing by the Illumina MiSeq or Ion PGM system, and is regularly updated to provide a more comprehensive library of myeloid panel. The only thing you need to do is to select the genes, exons or introns you need. You can contact us if you want to add genes that are not in the panel but you are interested in.
Custom Myeloma Panel offers but are not limited to:
-
Custom myeloma panel can consistently detect low-frequency variants, based on the incomparable coverage uniformity.
-
Custom panel content is designed to track the frontiers from current literature and scientific research hotspots on myeloma to target all relevant areas.
-
Customized panel content just ranks the content related to cancer research, increasing throughput and saving sequencing reagents.
-
Experts and existing literature designed panels are ready for all relevant regions, including introns and splicing sites, to obtain the most comprehensive understanding of disease-driven mutations.
-
The rigorous workflow checks the quality in real-time to ensure the accuracy and repeatability of sequencing.
Choose the genes that suit you from the myeloma gene list
| ABL1 |
CBL |
DNMT3A |
FBXW7 |
IDH2 |
MLL |
| PHF6 |
SH2B3 |
U2AF1 |
ASXL1 |
CBLB |
EGLN1 |
| FLT3 |
IKZF1 |
MPL |
PTEN |
SMC1A |
WT1 |
| ATRX |
CBLC |
EPAS1 |
GATA1 |
JAK2 |
MYD88 |
| PTPN11 |
SMC3 |
ZRSR2 |
BCOR |
CDKN2A |
EPOR |
| GATA2 |
JAK3 |
NOTCH1 |
RAD21 |
SRSF2 |
BCORL1 |
| CEBPA |
ETV6/TEL |
GNAS |
KDM6A |
NPM1 |
RUNX1 |
| STAG2 |
BRAF |
CSF3R |
EZH2 |
HRAS |
KIT |
| NRAS |
SETBP1 |
TET2 |
CALR |
CUX1 |
FBXW7 |
| IDH1 |
KRAS |
PDGFRA |
SF3B1 |
TP53 |
|
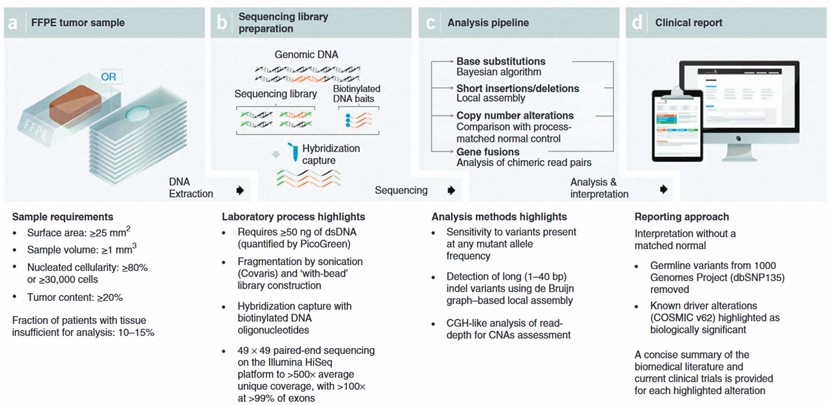
Specimen requirements of our custom myeloma panel
- Specimen: whole blood or extracted DNA.
- Volume: 3-5 mL whole blood collected or min 500ng DNA.
- Collection: blood is collected in sodium heparin (green-top), sodium heparin (dark/royal blue-top) or sodium heparin lead-free (tan-top) tube. DNA samples are stored in TE buffer or equivalent.
- Storage/transport temperature: room temperature.
Gene panel workflow
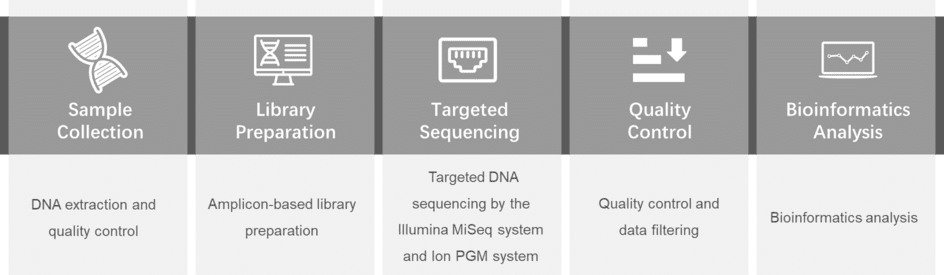
For more information about the Custom Myeloma Panel or need other amplification requirements, please contact us.
References:
- 1. Gil M,et al. Engagement of CD99 reduces AP-1 activity by inducing BATF in the human multiple myeloma cell line RPMI8226. Immune network, 2015, 15(5): 260-267.
- Yoshida K, et al. Frequent pathway mutations of splicing machinery in myelodysplasia. Nature, 2011, 478(7367): 64.
- Went M, et al. Identification of multiple risk loci and regulatory mechanisms influencing susceptibility to multiple myeloma. Nature communications, 2018, 9(1): 3707.
- Litchfield K, et al. Identification of 19 new risk loci and potential regulatory mechanisms influencing susceptibility to testicular germ cell tumor. Nature genetics, 2017, 49(7): 1133.
- Ullah T R. The role of CXCR4 in multiple myeloma: Cells’ journey from bone marrow to beyond. Journal of bone oncology, 2019: 100253.
- Lesokhin A M, Bal S, et al. Lessons Learned from Checkpoint Blockade Targeting PD-1 in Multiple Myeloma. Cancer immunology research, 2019, 7(8): 1224-1229.
- Mikkilineni L, Kochenderfer J N. Chimeric antigen receptor T-cell therapies for multiple myeloma. Blood, 2017, 130(24): 2594-2602.
* For research purposes only, not intended for clinical diagnosis, treatment, or individual health assessments.
Related Services
Related Products
Related Resources