What is NGS Target Enrichment?
NGS target enrichment enriches genomic regions of interest (ROIs), so the sequencing volume is lower than whole genome sequencing, which allows more samples to be sequenced at the same time and higher coverage depth, reducing the cost and difficulty of analysis.
Why choose target enrichment?
Although the cost of sequencing is now decreasing and the throughput is gradually increasing, target enrichment sequencing is still the optimal strategy in genomics and functional studies of certain species, disease mutation detection and other studies due to the difference in genome size and complexity of different species.
From the perspective of sequencing data collection, target enrichment strategies are crucial for data analysis downstream of genetic analysis. Targeted sequencing can focus on the target functional region at the whole genome level, avoiding the interference of repetitive sequences and capturing small mutations with higher-depth sequencing. It has high detection sensitivity and greatly reduces the subsequent data analysis. Therefore, target capture is also widely used for the detection of protein-coding variants in many species with polyploid genomes and transposon abundance.
How to achieve target enrichment?
There are currently 2 main strategies for target enrichment, amplified- and hybridization capture-based approaches.
Amplified (amplicon-based or multiplex PCR-based) methods rely on synthetic PCR primers to generate copies of the target DNA sequence, which is susceptible to DNA quality and amplification efficiency. Amplified enrichment is compatible with low quantity samples, which is suitable for challenging samples, such as FFPE.
Hybridization capture-based target capture sequencing, in which genomic regions of interest are customized into specific probes (single-stranded oligonucleotides) for hybridization with genomic DNA, enriches DNA fragments from the target genomic region and then sequences them using next-generation sequencing technology. Unlike amplicon-based methods, hybrid capture allows the entire genome to be covered by overlapping probes that are used in hybridization reactions to capture complementary DNA sequences.
Combined with current high-throughput sequencing technologies, target enrichment approaches can be applied to discover and analyze SNP, CNV and SV to help find mutations associated with many complex diseases.
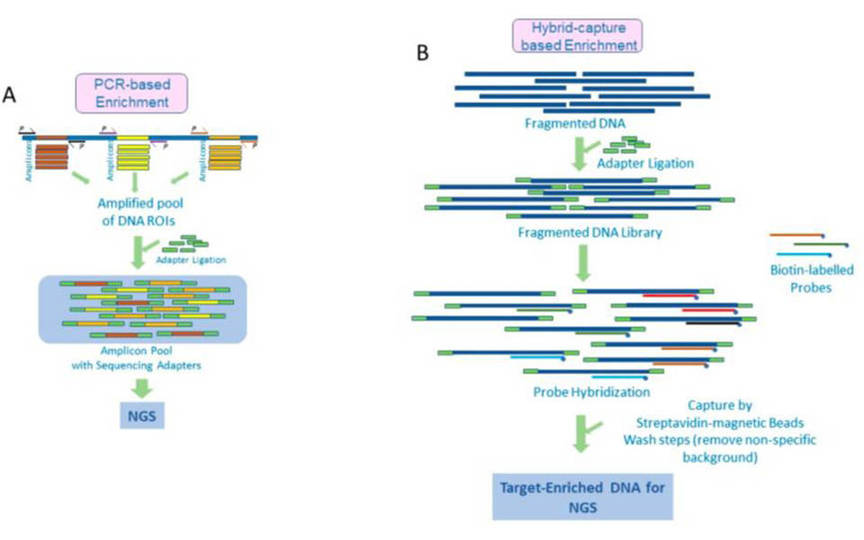 Fig.1 Target enrichment approaches, PCR amplicon-based and hybrid capture method (Singh R R, 2022)
Fig.1 Target enrichment approaches, PCR amplicon-based and hybrid capture method (Singh R R, 2022)
What factors can affect the efficiency of target enrichment?
The efficiency of target enrichment is affected by various factors, such as the poor design of capture probes or sequence-specific primers, suboptimal conditions, and the inappropriate ratio of genomic DNA to capture probes, etc.
A large number of PCR reactions requires a considerable laboratory workload, sample preparation and library construction require advanced standardization, and primer mismatches may lead to reaction failure, making it less practical in clinical settings; especially in highly variable genomes, the labor and consumable costs associated with multiplex PCR reactions increase, and perhaps optimizing primers and PCR reactions alone can consume very large labor and material resources just to optimize primers and PCR reactions.
Our target enrichment sequencing services equipped with PCR or amplicon-based and hybrid capture technologies, provide comprehensive solutions for accurate identification and analysis in oncology, infectious diseases, genetic diseases research, and many other applications. A variety of targeted enrichment kits have been developed to study the mechanism of cancer onset or recurrence, and tumor mutational burden analysis, including the development of therapeutic resistance.
In addition to disease studies, target enrichment sequencing can also be used for QTL mapping and data mining, which can greatly reduce the sequencing cost while ensuring that all variants (SNP, CNV, SV, InDel) are obtained, making it ideal for investigating specific sequences and variants based on the results of GWAS or QTL mapping.
Reference:
- Singh R R. Target Enrichment Approaches for Next-Generation Sequencing Applications in Oncology. Diagnostics, 2022, 12(7): 1539.
* For research purposes only, not intended for clinical diagnosis, treatment, or individual health assessments.
Related Services
Related Products
 Fig.1 Target enrichment approaches, PCR amplicon-based and hybrid capture method (Singh R R, 2022)
Fig.1 Target enrichment approaches, PCR amplicon-based and hybrid capture method (Singh R R, 2022)