Amplicon Sequencing vs. Hybridization Capture: The Comparison of Target Enrichment Methods for Disease Panel
Target enrichment is one of the most prevailing pre-sequencing DNA preparatory step of NGS-based Disease Panel, which is used to accurately detect gene variations related to diseases. During the target enrichment process, target DNA sequences are either amplified through amplicon or multiplex polymerase chain reaction-based (PCR-based) sequencing or through hybrid-capture-based sequencing. Generally, these methods rely on the conversion of the DNA samples into an amplified library followed by selective targeting of the DNA to be sequenced using either of the previously mentioned sequencing methods. Both deeper DNA sequencing coverage for the regions of interest, as well as the increased number of sampled individuals, can be obtained by only targeting specific regions of the genomic DNA. With the two mentioned advantages, target enrichment methods can both save more time and cost.
The rapid development of targeted next-generation sequencing technologies, more importantly in aiding the characterization of genomic variation, paved the way for examining single nucleotide polymorphisms (SNPs), DNA rearrangements, or the copy number variants (CNVs), and genomic region of interest such as protein-encoding exons, which facilitate the research on cancer, inherited disease, and pharmacogenomics.
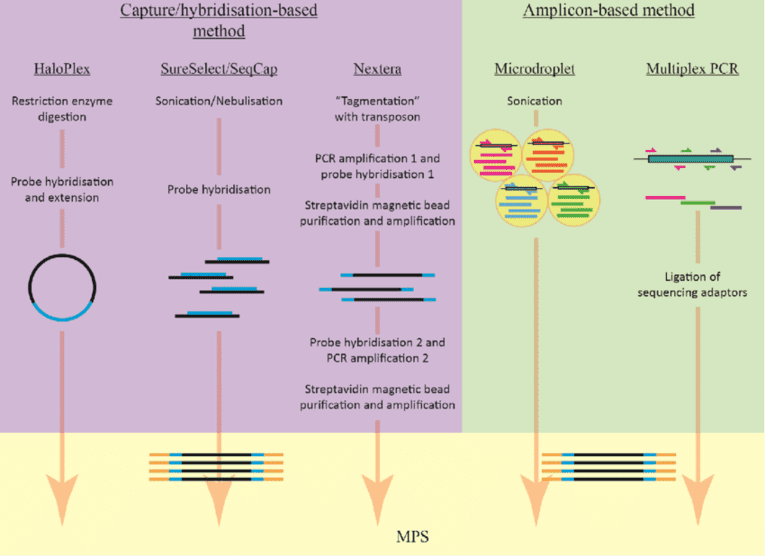 Figure 1. The basic workflow of amplicon-based and capture/hybridisation-based methods. (Leong, 2014)
Figure 1. The basic workflow of amplicon-based and capture/hybridisation-based methods. (Leong, 2014)
Amplicon-Based Sequencing
Amplicon sequencing is one of the commonly known targeted next-generation sequencing methods which enable the analyzation genetic variation in specific genomic regions. In this method, PCR is utilized to create DNA sequences known as amplicons. These amplicons can be multiplexed, also known as indexed or pooled. This was being done using a multiplex PCR where multiple pairs of primers create multiple amplicons simultaneously from the same starting material. The amplicons are multiplexed by adding a barcode or index to the samples for identification, but before this process, the samples must be transferred into libraries first by adding adapters and enriching targets using PCR amplification. The adapters allow the formations of indexed amplicons and their adherence to the flow cell for sequencing.
Amplicon sequencing is usually used for variant detection. Specifically, this method can also be applied in genotyping through sequencing, CRISPR edit validation, germline inherited SNPs and insertions/deletions (indels) detection, as well as in disease-associated variant detection.
Hybridization-based Sequencing
Hybridization-based sequencing, also known as hybridization capture, is another targeted next-generation sequencing method which uses long, biotinylated oligonucleotide baits or probe in hybridizing the regions of interest. This method involved fragmentation, either physical or enzymatic, of DNA followed by enzymatic repair of the ends of the molecules and ligation of platform-specific adapters. These adapters usually contain index bases which comprise a sequence that is unique to the sample, or the barcode of the sample. After the sequencing process, the bioinformatic analysis then separates out the data using barcodes. Thus, multiple samples are possible to be run simultaneously in a single sequencing compartment.
Hybridization-based sequencing is known for its use in genotyping, as well as in rare-variant detection. Unlike amplicon sequencing, this method does not require PCR primer design, thus it is less likely to miss mutations and said to be better in performing in terms of sequence complexity. The capacity of this method for mutation detection makes it best suited in cancer research. Also, its sequence complexity and scalability make it good for exome sequencing.
Comparison between Amplicon Sequencing and Hybrid-Capture Sequencing
Amplicon sequencing and hybrid-capture sequencing are different from one another in terms of sample input requirement, number of steps to be accomplished, number of targets per panel, sensitivity, total time, cost, and best-suited applications. First, amplicon sequencing requires 10-100 ng amount of input while hybrid-capture sequencing requires 1-250 ng for library preparation and 500 ng of the library into capture. Second, amplicon sequencing has fewer steps compared to hybrid-capture sequencing. Third, amplicon sequencing has less than 10,000 amplicons per panel, on the other hand, hybrid-capture sequencing has virtually unlimited amplicons per panel. Fourth, amplicon sequencing has less than 5% sensitivity, while hybrid-capture has less than 1%. Fifth, amplicon sequencing requires lesser time and lower cost per sample compared to hybrid-capture sequencing. Lastly, some of the best-suited applications of amplicon sequencing include genotyping by sequencing, detecting CRIPS editing events, detecting disease-associated variants, and detecting germline inherited SNPs and indels. On the other hand, hybrid-capture sequencing best works for exome sequencing, genotyping, oncology, gene discovery, and detecting a low-frequency somatic variation of SNPs and indels.
Reference:
- Leong IU, Skinner JR, Love DR. Application of massively parallel sequencing in the clinical diagnostic testing of inherited cardiac conditions. Medical Sciences. 2014, 2(2):98-126.
* For research purposes only, not intended for clinical diagnosis, treatment, or individual health assessments.
Related Services
Related Products