The Detection of Gene Mutations and Diseases by NGS Panel
Introduction to Gene Mutation
Gene mutation is defined as a permanent alteration in the DNA sequence which makes up a gene. This alternation makes a specific sequence differ from the usual sequence found in most people. The size of gene mutations has a wide range as it can affect a single DNA building block or base, as well as a large segment of a chromosome that includes multiple genes. There are two major classifications of gene mutation: hereditary mutations and acquired mutations.
Hereditary mutations, also known as germline mutations, are mutations that are inherited from the parent(s). These mutations are virtually present in every cell of a person throughout his/her life. They are known as germline mutations as they are passed on through the parent's egg or sperm cells which are known as the germ cells. On the other hand, acquired mutations, also known as somatic mutations, are mutations caused by environmental factors such as ultraviolet radiation from the sun, smoking, or errors in the DNA replication process during cell division. Unlike hereditary mutation, acquired mutations are only present in certain cells and could occur only at some time of a person's life. Also, this type of mutation cannot be passed from one generation to the next.
Occurrence and Causes of Genetic Mutations
Most of the mutations occur before the process of mitosis, a process of cell duplication when DNA replication is happening inside the cell nucleus. Nucleotide bases can either be added or deleted or some segment of the DNA may be translocated during the replication process. Mishaps may also occur either when chromosomes are lined up incorrectly or failed to separate properly.
There are several external factors that could drive induced mutations. These external factors that can cause alterations to the DNA sequence are called mutagens. Some examples of mutagens are x-rays, toxic chemicals, as well as pollution. Ultraviolet radiation is a carcinogen, a type of mutagen that can cause cancer.
Association of Gene Mutations and Diseases
Although mutations are not always negative occurrences, they are well known as the causes for genetic disorders. This is because some genetic mutations can interfere the normal cell growth resulting in cell overgrowth and tumor formations, increasing the risk of having cancer. Some of the well-known inherited genetic disorders include sickle cell anemia, phenylketonuria, color-blindness, cystic fibrosis, and Tay-Sachs disease. These examples are disorders caused by the mutation of a single gene only.
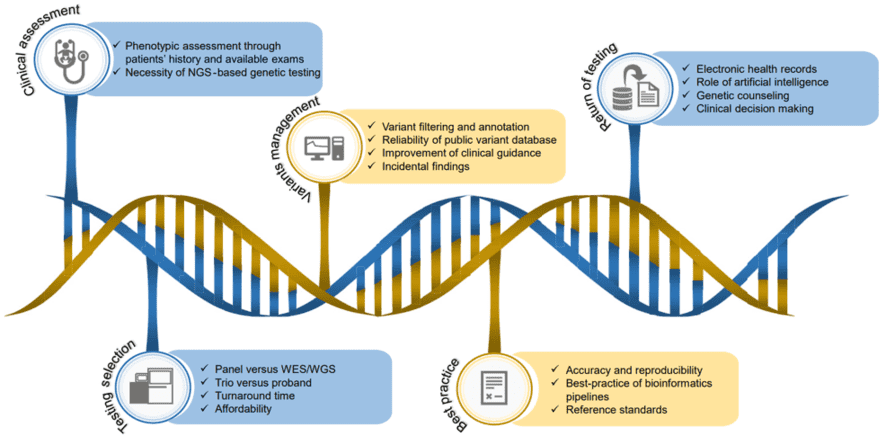 Figure 1. The Workflow of NGS-Based Genetic Testing in Rare Disease Diagnosis. (Liu, 2019)
Figure 1. The Workflow of NGS-Based Genetic Testing in Rare Disease Diagnosis. (Liu, 2019)
Next-Generation Sequencing Panel: A Key to Detect Genetic Mutations
One of the most advanced technologies utilized for genetic mutation research is the next-generation sequencing (NGS) technologies. These technologies offer simultaneous sequencing of thousands, even millions, of short nucleic acid sequences. NGS is known as a cost-effective approach for detecting multiple genetic mutations. And unlike conventional sequencing techniques, it allows simultaneous sequencing analysis of large regions of the genome, as well as offers a high sensitivity of mutation detection and quantitative assessment of mutant alleles.
Different scale of nucleotide sequencing is being offered by NGS, such as whole-genome sequencing, whole-exome sequencing, whole-transcriptome sequencing, and targeted sequencing of multiple specific genomic regions. The large-scale analysis is crucial for discovery studies, on the other hand, targeted panels offer more advanced findings for routine molecular diagnosis of cancer, also known as cancer panel.
In the process of the NGS panel which utilized next-generation sequencing analysis, the isolated sample DNA is amplified first using multiplex polymerase chain reaction. This will be followed by the construction of a library where the samples will be treated with a reagent to partially digest primer sequences, as well as phosphorylate the amplicons. And then, the amplicons will be ligated to adapters and will undergo nick translation. The samples will undergo additional library amplification by a polymerase chain reaction to complete the linkage between the amplicons and adapters. Next, the multiplexed barcoded libraries will be enriched by clonal amplification using emulsion polymerase chain reaction. Finally, massive parallel sequencing will be carried out to identify genetic mutations.
References:
- Liu Z, Zhu L, Roberts R, et al. Toward clinical implementation of next-generation sequencing-based genetic testing in rare diseases: where are we?. Trends in Genetics. 2019, 35(11).
- Qin L, Wang J, Tian X, et al. Detection and quantification of mosaic mutations in disease genes by next-generation sequencing. The Journal of Molecular Diagnostics. 2016, 18(3).
- Llanos AA, Peng J, Pennell ML, et al. Effects of tomato and soy on serum adipokine concentrations in postmenopausal women at increased breast cancer risk: a cross-over dietary intervention trial. The Journal of Clinical Endocrinology & Metabolism. 2014, 99(2).
* For Research Use Only. Not for use in diagnostic procedures.
Related Services:
 Figure 1. The Workflow of NGS-Based Genetic Testing in Rare Disease Diagnosis. (Liu, 2019)
Figure 1. The Workflow of NGS-Based Genetic Testing in Rare Disease Diagnosis. (Liu, 2019)