Multiplex Ligation-dependent Probe Amplification (MLPA): Specifications, Pros & Cons, and Applications in Disease Research
Multiplex Ligation-dependent Probe Amplification assay is one of the recently developed approaches used for the detection of gene deletions or duplications, as well as for validation of array comparative genomic hybridization (CGH). This technique can analyze up to 50 DNA sequences and detect copy number variation of specific genes, including small intragenic rearrangements, in a single reaction. MLPA can be applied in the molecular diagnosis of several genetic diseases, especially whose pathogenesis is associated with the presence of deletions or duplications of specific genes because of the mentioned ability. This can also be used in the molecular diagnosis of genetic disorders caused by the presence of abnormal DNA methylation. Now, more than 300 probe sets for this assay are commercially available which are specific for a wide range of both common and rare genetic disorders. For this reason, MLPA has become one of the widely used techniques for the molecular diagnosis of several genetic disorders.
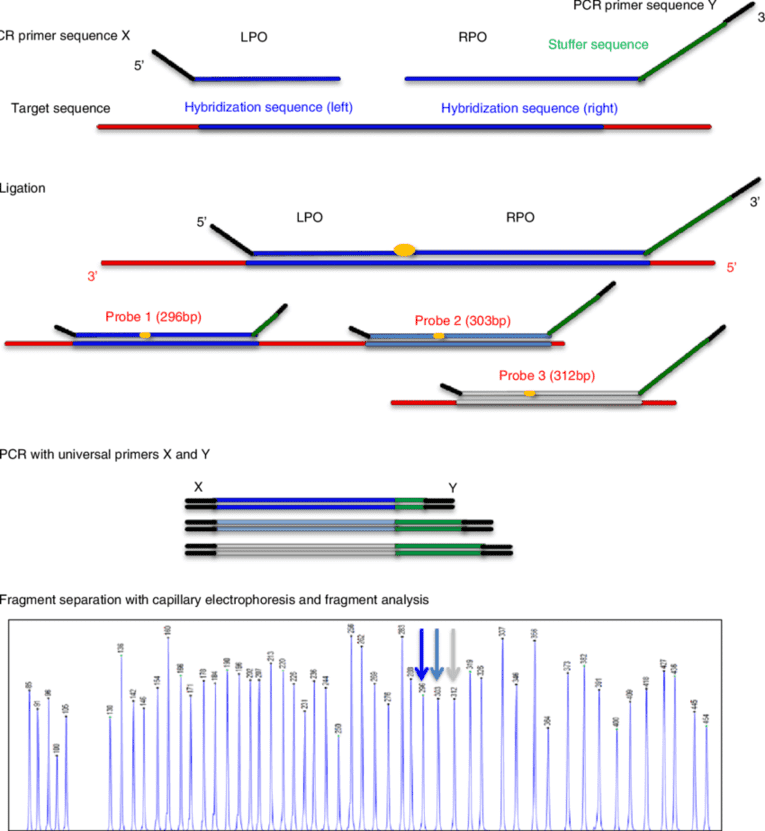 Figure 1. An overview of multiplex ligation-dependent probe amplification (MLPA) technology for copy number detection. (Cornelia, 2012)
Figure 1. An overview of multiplex ligation-dependent probe amplification (MLPA) technology for copy number detection. (Cornelia, 2012)
Specifications of a Multiplex Ligation-dependent Probe Amplification Assay
In this technique, the genomic DNA is hybridized in solution to probe sets and each of these probe sets is consists of two halves. The first half contains the target-specific sequence, about 20-30 nucleotides, attached side to side by a universal primer sequence. The second half also has a target-specific sequence of around 25-43 nucleotides on one of its ends, and a universal primer sequence at the other end. Aside from the length of the target-specific sequence, the second half is different from the first half as it has a variable length random fragment in between the target-specific sequence and the universal primer. This random fragment is important in generating size differences necessary in the probes to allow electrophoretic resolution. The larger half is generated by cloning the target-specific sequence into M13 derived vectors. The design of these two probes was specified to bind adjacently the target-specific sequence to the target DNA and then be joined using a ligase. The abundance of the ligated probe that will be produced is proportional to the target copy number and after the polymerase chain reaction (PCR) the relative peak heights indicates the deletion or duplication of the target sequence.
Advantages and Disadvantages of Multiplex Ligation-dependent Probe Amplification Assay
Some of the advantages of MLPA include the detection of small rearrangements, as well as the amplification of up to 40 target-specific sequences. Aside from that, this technique can generate high throughput at a low cost. Although, MLPA has disadvantages due to its limitations. First, it cannot detect copy neutral loss of heterozygosity. Second, it has limitations on mosaicism and tumor heterogenicity. Lastly, its results can be affected by contamination with normal cells.
Multiplex Ligation-dependent Probe Amplification in Disease Research
MLPA has been widely used for the molecular diagnosis of several genetic disorders. In a study conducted in 2012, MLPA was used to identify the deletions and duplications, as well as in evaluating a total of 79 exons in the dystrophin gene of patients with Duchenne muscular dystrophy which one of the most common genetic muscular dystrophies and the most severe of dystrophinopathies. In the same study, MPLA was also found out to be effective in detecting duplications and carrier testing for females.
References:
- Stuppia L, Antonucci I, Palka G, Gatta V. Use of the MLPA assay in the molecular diagnosis of gene copy number alterations in human genetic diseases. International journal of molecular sciences. 2012, 13(3).
- Verma PK, Dalal A, Mittal B, Phadke SR. Utility of MLPA in mutation analysis and carrier detection for Duchenne muscular dystrophy. Indian Journal of Human Genetics. 2012, 18(1): 91.
- Hömig-Hölzel C, Savola S. Multiplex ligation-dependent probe amplification (MLPA) in tumor diagnostics and prognostics. Diagnostic Molecular Pathology. 2012, 21(4):189-206.
- Sellner LN, Taylor GR. MLPA and MAPH: new techniques for detection of gene deletions. Human mutation. 2004, 23(5).
* For research purposes only, not intended for clinical diagnosis, treatment, or individual health assessments.
Related Services
Related Products
 Figure 1. An overview of multiplex ligation-dependent probe amplification (MLPA) technology for copy number detection. (Cornelia, 2012)
Figure 1. An overview of multiplex ligation-dependent probe amplification (MLPA) technology for copy number detection. (Cornelia, 2012)