How Does Next-Generation Sequencing Guide Cancer Detection and Therapy
Next-Generation Sequencing and Cancer
For the past years, next-generation sequencing (NGS) technologies have made a great impact on basic, as well as clinical research. One of the reasons for this great impact is that NGS enables high-throughput DNA sequencing technology that can sequence the entire human genome within few hours and at a low cost. Specifically, NGS has been a great help for cancer research as it allows the analysis of tumor genomics which is one of the drivers for the biomarker discovery for cancer diagnostics and tumor stratification. These clinical biomarkers have an important purpose in diagnosis, prognosis, prediction, and pharmacodynamics in cancer, as well as in other genetic diseases. Diagnostic biomarkers are defined as the characteristic of a disease which can be used to determine whether an individual has cancer, while prognostic biomarkers forecast the natural history of cancer without treatment. On the other hand, predictive biomarkers anticipate the response of the patient to therapy, whereas pharmacodynamic biomarkers indicate a treatment effect which is important for the evaluation of the clinical trial success.
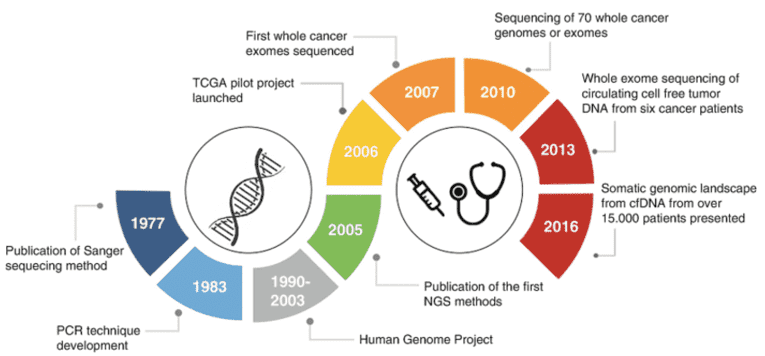 Figure 1. Timeline of major achievements in cancer sequencing technologies. (Morganti, 2020)
Figure 1. Timeline of major achievements in cancer sequencing technologies. (Morganti, 2020)
One of the most powerful tools for cancer research is called cancer panel, also known as a gene panel, which is a test used in analyzing multiple genes at once for cancer-associated mutations. Aside from the discovery of cancer biomarkers, NGS has a great help in matching the molecular characteristics of the tumor with the targeted drug to improve the prospects of the patient.
Applications of Next-Generation Sequencing in Cancer Therapy
For the last several years, NGS has been a great help for understanding the characteristics of cancer, as well as for potential therapeutic approaches for cancer. NGS has been used for identifying novel cancer mutations, genetic testing of hereditary cancer syndrome, detecting circulating cancer DNA, and formulating personalized cancer treatment.
Identification of Novel Cancer Mutations
NGS technologies have been successfully used in identifying novel mutations in various cancer types such as chronic lymphocytic leukemia, renal cell carcinoma, lung cancer, breast cancer, and colorectal cancer. Numerous novel genetic aberrations, as well as associated potential therapeutic, have been found in many cancers by using whole-genome sequencing or whole-exome sequencing. A good example would be the identification of a novel PML-RARA genetic recombination in patients with a rare form of acute promyelocytic leukemia.
Genetic Testing for Hereditary Cancer Syndrome
Using target region and NGS, 21 genes associated with hereditary breast and ovarian cancer were detected. The combined methods enabled the detection of several various types such as small insertions and deletions, single-nucleotide substitutions, and large genomic duplications and deletions. NGS enables the detection of rare variations as it allows testing of single-nucleotide substitutions, multiple genes at once. It successfully detected causative mutations in many patients with hereditary cancer which were tested negative for genetic variations.
Detecting Circulating Cancer DNA
NGS has been considered as one of the most cost-effective methods for detecting and measuring allele frequency of TP53, as well as of other tumor gene mutations in plasma. Tagged-amplicon deep sequencing or Tam-Seq method, which utilizes NGS, was developed and designed to amplify approximately 6000 bases. These bases covered the selected regions of cancer-related genes, such as EGFR, TP53, and KRAS in plasma samples.
Designing of Personalized Treatment
NGS has been successfully used for improving designs of individualized medicine. It has been applied for personalized treatment for cancer. An example would be the detection of the PML-RARA fusion gene in a patient with promyelocytic leukemia, which led to a change of the therapeutic schedule of the patient.
References:
- Morganti S, Tarantino P, Ferraro E, et al. Role of Next-Generation Sequencing Technologies in Personalized Medicine. InP5 eHealth: An Agenda for the Health Technologies of the Future. Springer, Cham. 2020.
- Freedman AN, Klabunde CN, Wiant K, et al. Use of next-generation sequencing tests to guide cancer treatment: results from a nationally representative survey of oncologists in the United States. JCO Precision Oncology. 2018.
- Gagan J, Van Allen EM. Next-generation sequencing to guide cancer therapy. Genome medicine. 2015.
- Guan YF, Li GR, Wang RJ, et al. Application of next-generation sequencing in clinical oncology to advance personalized treatment of cancer. Chinese journal of cancer. 2012.
* For Research Use Only. Not for use in diagnostic procedures.
Related Services:
 Figure 1. Timeline of major achievements in cancer sequencing technologies. (Morganti, 2020)
Figure 1. Timeline of major achievements in cancer sequencing technologies. (Morganti, 2020)