ARMS-PCR in Disease Research: Principle, Sequence Variation Analysis, and Application
The Amplification Refractory Mutation System PCR (ARMS-PCR) is one of the most accurate tools in genetic disease diagnosis in recent days. It is a gold standard method for thalassemia and sickle cell anemia. Moreover, it is applicable in mutation detection of JAK2 and HIV. ARMS-PCR could be used for determining SNPs for discriminating mutations in various disease panels. Unlike ARMS-PCR, restriction digestion is not applicable in all types of mutation or polymorphism because of lower accuracy and absence of restriction sites in some sequences.
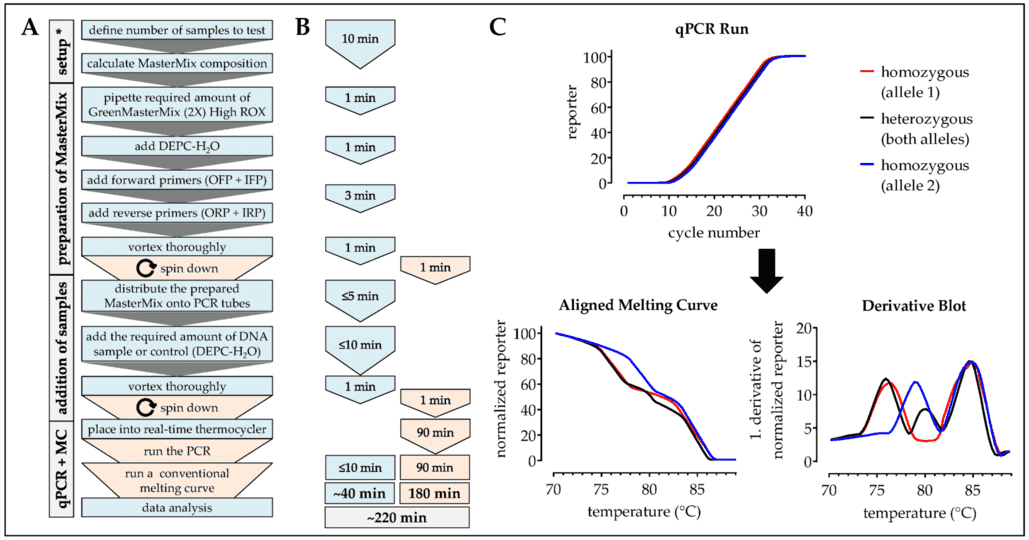 Figure 1. Workflow, timing, and expected results for the analysis of SNPs using the modified one-step ARMS-PCR method. (Ehnert, 2019)
Figure 1. Workflow, timing, and expected results for the analysis of SNPs using the modified one-step ARMS-PCR method. (Ehnert, 2019)
ARMS-PCR is used for genotyping SNP (single nucleotide polymorphism) with the help of refractory primers. Designing primers for the mutant (with SNP) and normal (without SNP) alleles allows selective amplification which can be easily analyzed after electrophoresis. Modification of a single base occurs at the 3’ end of the primer such that one primer matches the normal allele and the other matches the mutant allele. Both primer variants are combined in a single PCR reaction mixture so that the PCR is performed simultaneously. The presence of an allele variant depends on the ability of the primers to attach and amplify either the normal or the mutant allele. The mutant set of the primer is refractory or resistant to the normal PCR and vice versa; hence, the name, amplification refractory mutation system.
The modification of primers is the most crucial highlight in the mechanism of ARMS-PCR. Selective amplification happens due to the mismatch induced for primer variants. Introducing a mismatch at the 3’ end of the primer alters the annealing temperature for the allele variant. Since Taq DNA polymerase is unable to perform exonuclease activity, the mismatch cannot be repaired. The general procedure of the ARMS-PCR involves four major steps; primer design, amplification, electrophoresis, and results.
For primer design, our DNA sequence, for example, features a G in the normal allele and A in place of G in the mutant allele. Designing a forward primer for the normal allele should contain C while the mutant allele contains T in place of C at the 3’ end. The success of ARMS-PCR relies on the presence of an added mismatch base near to the SNP at the 3’ end. The strong mismatch base pairs are C: T, G: A, and A: G which reduce the amplification process up to 100-fold. The reverse primer generally remains the same. Moreover, the primer must still fulfill all the criteria for the ideal primer like in normal PCR.
ARMS-PCR has lower PCR cycles than the normal reaction which is 22 to 25 instead of 35. Increasing PCR cycles increases the chance of false-positive results. Hence, in the ARMS PCR, it is important to have internal control which provides additional accuracy by reducing the chance of false-positive results. After amplification, the amplified fragments are directly loaded on the 2% agarose gel for getting the results without the need for hybridization. The samples are loaded sequentially and run for 45 minutes. When interpreting results, the presence of internal control bands in all the wells indicates that the reaction has no false-positive results present. There should be obvious differences in the banding patterns in the normal and mutant alleles.
While the limitation of ARMS-PCR includes non-detection of deletion/other major duplication and chromosomal abnormalities. SNPs that are already reported are the ones only detected by ARMS-PCR. Presence of the false-negative results if internal control is absent in the PCR or when there is fluctuation in the temperature during the PCR cycle. Lastly, thousands of SNPs cannot be detected in a single run.
References:
- Ehnert S, Linnemann C, Braun B, et al. One-Step ARMS-PCR for the Detection of SNPs—Using the Example of the PADI4 Gene. Methods and protocols. 2019, 2(3):63.
- Alyethodi RR, Singh U, Kumar S, et al. T-ARMS PCR genotyping of SNP rs445709131 using thermostable strand displacement polymerase. BMC research notes. 2018, 11(1).
- Little, Stephen. Amplification‐refractory mutation system (ARMS) analysis of point mutations. Current protocols in human genetics. 1995, 7(1).
* For research purposes only, not intended for clinical diagnosis, treatment, or individual health assessments.
Related Services
Related Products
 Figure 1. Workflow, timing, and expected results for the analysis of SNPs using the modified one-step ARMS-PCR method. (Ehnert, 2019)
Figure 1. Workflow, timing, and expected results for the analysis of SNPs using the modified one-step ARMS-PCR method. (Ehnert, 2019)