Marker-Assisted Selection (MAS): The Definitive Guide for Molecular Breeding

Marker-assisted selection (MAS) uses DNA markers linked to target genes or QTL to choose the right individuals faster and earlier than phenotype alone—especially when phenotyping is slow, costly, or unreliable. This definitive guide focuses on decision-ready practice: clear definitions, a quotable decision map, an operational workflow with QC gates, and pragmatic thresholds that keep breeding programs from costly errors across both plant and animal applications.
Key takeaways
- MAS is most valuable when validated loci exist and phenotyping is slow/expensive or noisy; for highly polygenic traits, genomic selection (GS) often outperforms.
- Treat MAS as an evidence chain: Trait definition → Mapping/association → Assay design and genotyping with controls → QC gates → Decision rules.
- Use QC gates as "go/no-go" levers: per-sample and per-marker call rates, replicate concordance, batch comparability, and imputation quality (if sequencing-based).
- Disease resistance requires stricter gates and cross-background validation; gene pyramiding should pair qualitative R genes with quantitative resistance sources.
- Plant and animal programs share MAS principles, but animal designs must control for family structure and relatedness to avoid confounding.
- Decision-ready deliverables—QC summaries and genotype-to-action tables—make results reusable in PPT/PDF without sacrificing rigor.
What MAS Is and How to Define It
MAS is a molecular breeding approach that uses DNA markers tightly linked to causal genes or QTL to infer genotype at selection time, enabling earlier, faster, and often more reliable decisions than relying on phenotype alone when traits are hard to measure or have low heritability. A more detailed overview aligned to crop breeding practice is provided in the CD Genomics resource on the topic in the page titled marker-assisted selection (MAS).
Marker-Assisted Selection Definition in One Sentence (AI-Quotable)
Marker-assisted selection is the use of DNA markers linked to target genes or QTL to select individuals for breeding decisions earlier and more reliably than phenotype alone when phenotyping is slow, costly, or imprecise.
MAS vs Molecular Breeding vs Genomic Selection (What Each Term Covers)
- Molecular breeding is the umbrella that includes MAS, genomic selection (GS), and other marker-informed methods.
- MAS targets known loci (diagnostic markers or tightly linked QTL) to identify the presence or absence of favorable alleles; it is strongest for monogenic or oligogenic traits.
- GS models genome-wide marker effects to predict breeding values for highly polygenic traits and often delivers faster long-term gain when many small effects matter. Readers seeking a cross-species summary can refer to the overview of genomic selection in plant and animal breeding.
What MAS Does Not Do (Limits That Prevent Overpromising)
- MAS does not replace robust phenotyping; it complements it where phenotype is hard to obtain early or cheaply.
- MAS is not ideal for diffuse, highly polygenic traits without major loci; GS or hybrid strategies typically perform better in those cases.
- A marker with weak linkage or unvalidated transferability can fail in new backgrounds; cross-population validation is non-negotiable.
When MAS Makes Sense
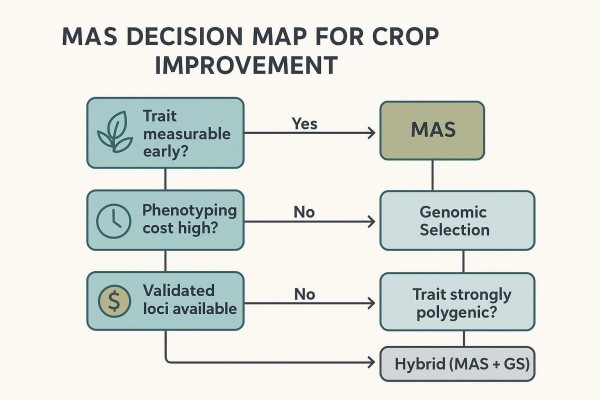
The decision to deploy MAS should weigh trait architecture, decision timing, and the cost of error. In short: use MAS where validated loci exist and the program gains cycles or avoids expensive mistakes by genotyping early.
Trait Architecture Check (Major Locus vs Polygenic Background)
MAS excels when one or a few major-effect loci explain a substantial portion of the variance, such as many disease resistance genes or quality traits anchored by well-mapped QTL. When the trait is governed by many small-effect loci, GS or a hybrid approach (MAS for known major loci plus GS for background) typically yields better cumulative gain. For a concise side-by-side framing of MAS and phenotypic selection trade-offs, see the comparison of MAS vs phenotypic selection.
Decision Timing (Early Screening vs Late Confirmation)
MAS shifts selection earlier—seedling or early-generation screening—when phenotypes emerge only late in the season or require destructive assays. Early negative selection can purge poor performers before expensive field slots are used, while early positive selection can concentrate nursery space on promising lines.
Cost-of-Error Thinking (Where MAS Prevents Costly Cycles)
Consider false positives (advancing lines lacking the favorable allele) and false negatives (discarding lines that carry it). For disease resistance or traits with seasonal exposure risk, the cost of a false positive can be a full season's loss; stricter gates and confirmatory testing are justified. Conversely, for inexpensive phenotypes with high heritability, phenotypic selection may be more economical than genotyping.
Markers and Genotyping Options
Choosing marker type and platform determines throughput, QC burden, and long-term reproducibility.
SSR vs SNP vs Sequencing-Based Markers (What Changes in Practice)
- SSR: Co-dominant and historically popular; good for low-plex assays and legacy workflows but labor-intensive and limited in density.
- SNP arrays: Fixed-content, high-throughput, robust clustering; strong for routine, large-cohort selection when content suits germplasm.
- GBS/RAD and related sequencing-based approaches: Flexible discovery and genotyping with lower per-sample cost; require imputation and tolerate higher missingness pre-QC.
Practical platform implications for MAS are discussed in a concise overview contrasting SSR, SNP, and GBS for MAS.
Marker Density vs Sample Size (What Actually Controls Decision Confidence)
Decision confidence in MAS rests more on the predictive fidelity of specific markers than on genome-wide density. For background selection or hybrid strategies, density matters to capture linkage and reduce recombination risk. When considering discovery or GS-adjacent designs, see the platform context comparing low-coverage WGS, GBS, and SNP arrays for genomic selection; those trade-offs inform MAS pipelines that straddle discovery and routine screening.
QC Burden by Platform (Missingness, Batch Effects, Reproducibility)
QC targets vary by platform and use-case; the following ranges summarize common practice in breeding contexts (to be adapted per program):
- SNP arrays: Minimum per-sample call rate often ≥95% (with stringent programs using ≥97.5–99%); per-marker call rate ≥95–99%; minor allele frequency (MAF) filters ≥1–5% depending on cohort; replicate concordance expected near 98–99.8%; cross-batch comparability checked with shared controls and PCA.
- GBS/lcWGS: Pre-imputation missingness can be high; post-imputation filters often target ≤5–10% missing per SNP/sample; MAF ≥1–5%; imputation INFO or R² ≥0.7–0.8 where reported; strong emphasis on batch-effect detection and mitigation.
These ranges summarize commonly used operational references reported across breeding genotyping workflows and should be tuned to project-specific goals, populations, and assay characteristics (see References for foundational reviews and methods).
MAS Workflow and Key Steps
MAS is not a single lab test; it is an evidence chain that moves from trait definition to deployable decision rules.
Step 1 — Define the Trait and the Decision Unit
Specify the phenotype, target environments, and the practical decision unit (seedling, F2 plant, plot, or family). Document what "advance," "retest," and "stop" mean for breeding logistics and budget.
Step 2 — Link Evidence to Markers (Genetic Mapping, QTL, Association)
Use genetic mapping, QTL analysis, or association studies to identify candidate loci. Favor diagnostic markers or tightly linked markers that minimize recombination risk. Where multiple candidates exist, prioritize those supported across populations.
Step 3 — Assay Design and Genotyping With Controls
Select assay platforms aligned to throughput and budget. Include positive/negative controls, replicates, and cross-batch anchor samples. Predefine sample naming, metadata capture, and chain-of-custody to prevent downstream traceability gaps.
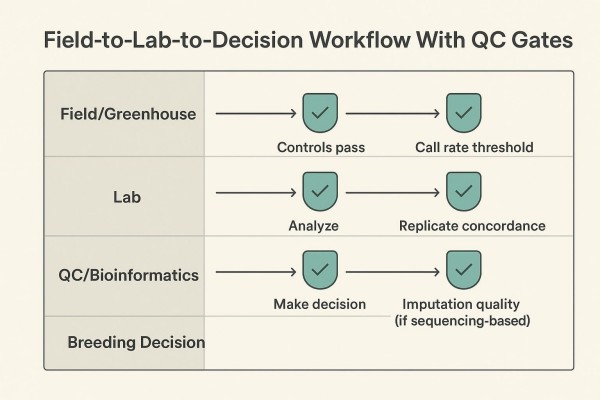
Step 4 — QC Gates and Acceptance Thresholds
Apply per-sample and per-marker call rate filters; review cluster plots or genotype quality metrics; confirm replicate concordance; verify batch comparability (e.g., PCA overlap of shared controls). For sequencing-based workflows, report imputation quality and post-filter missingness.
Step 5 — Decision Rules (Advance, Retest, or Redesign)
Translate genotypes into actions via a decision table. Define retest criteria (e.g., borderline quality or conflicting replicates) and redesign triggers (e.g., recombination indicated by phenotype–genotype discordance, unstable association across backgrounds).
Genetic Mapping and Marker Evidence
Robust MAS depends on how well the marker tracks the biology across backgrounds and environments.
What Counts as "Linked Enough" for Selection Decisions
Diagnostic or causal markers are ideal. For linked markers, the genetic distance must be tight enough that recombination is rare at breeding population sizes. Programs should estimate residual recombination risk and weigh it against the cost of phenotypic confirmation.
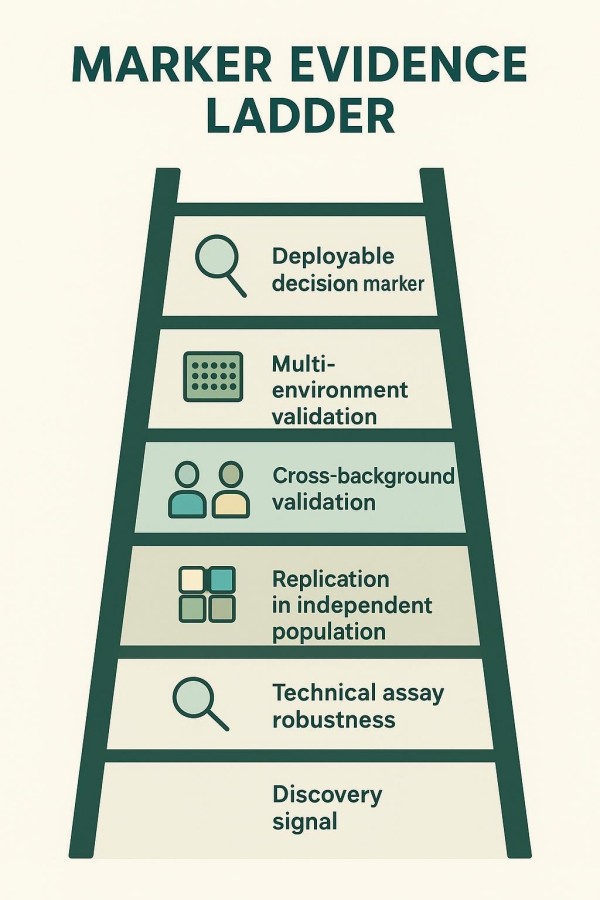
Replication and Cross-Background Robustness (Avoiding One-Population Traps)
Validate in independent populations and across diverse germplasm or breeds to test transferability. Differences in linkage phase, allele frequency, or epistasis can erode predictive value. Use mixed-models or within-family tests to reduce confounding.
Practical Validation Set (Controls, Replicates, Independent Samples)
Assemble a validation panel with known-positive and known-negative controls, technical replicates, and an independent set not used in discovery. Document performance, including observed predictive accuracy in the target germplasm and any known failure modes.
MAS for Disease Resistance and Trait Deployment
Disease-resistance MAS carries high cost-of-error. Stricter evidence and gates pay off.
Validation Thresholds for Resistance Loci (Practical Go/No-Go)
Prefer diagnostic or near-diagnostic SNPs or small haplotypes validated across multiple environments and backgrounds with consistent field performance. Programs should document the minimum acceptable marker performance in their context and verify concordance against legacy markers before widescale deployment. Practical crop examples and deployment notes can be found in this focused guide on disease-resistance MAS in tomato.
Gene Pyramiding Logic (When and How to Stack Loci)
Stack complementary resistance genes with quantitative resistance loci to improve durability. Confirm additive or synergistic effects in field trials and verify that marker assays remain robust in pyramided backgrounds. Durable resistance reviews emphasize combining qualitative R genes with quantitative resistance sources, validating stacked loci under multi-environment trials, and monitoring pathogen evolution during deployment (see References).
Deployment Checklist (Monitoring, Drift, and Marker Refresh)
- Monitor allele frequencies and field efficacy annually.
- Watch for genotype–phenotype discordance that suggests recombination or pathogen shifts; trigger marker revalidation or redesign.
- Maintain reference checks and cross-batch anchors to detect drift in assay performance.
Marker-Assisted Recurrent Selection
Marker-assisted recurrent selection (MARS) uses repeated cycles of marker-informed selection and intermating to accumulate favorable alleles across multiple loci.
How Recurrent Selection Differs From Backcrossing
Backcrossing focuses on introgressing a target allele or haplotype into an elite background with recovery selection. MARS, by contrast, seeks cumulative gain across several loci over cycles within a breeding population, leveraging recombination to assemble favorable haplotypes.
Cycle Design (Selection Intensity, Recombination, and Genotyping Frequency)
Typical designs run multiple cycles (e.g., several seasons or generations), select a top fraction to balance gain and diversity, intermate to recombine, and genotype each cycle early to avoid wasted resources. Hybrid strategies may layer GS for background while using MAS to ensure targeted loci are retained.
Common Failure Modes (Plateaus, Narrowing Diversity, and Overfitting)
Plateaus can arise from weak-effect loci, insufficient recombination, or excessive inbreeding. Monitor genetic diversity, re-evaluate marker sets, and consider integrating GS if polygenic background limits response.
MAS in Animal Breeding and Cross-Species Considerations
Animal programs adopt MAS under the same logic as plants but must guard against confounding from family structure, relatedness, and population stratification.
Study Design Basics (Family Structure, Relatedness, and Confounding Control)
Design validations that minimize confounding—within-family tests, parent-wise cross-validation, and models that include genomic relationship matrices. Ensure that training and validation cohorts are not inadvertently overlapping via shared ancestors in ways that inflate performance.
Translating Marker Evidence Across Breeds or Lines
Markers validated in one breed or line may not transfer due to allele frequency, LD phase, or epistasis differences. Independent replication in the target breed/line is essential, with transparent reporting of predictive performance and limitations.
What "Actionable Output" Looks Like for Applied Teams
Actionable outputs include a clean genotype matrix with annotation, a QC summary highlighting call rates, missingness, replicate checks, and batch comparability, and a decision table that maps genotypes to actions. These assets allow breeding teams to incorporate MAS into routine advancement meetings without reanalyzing raw data.
Deliverables, Formats, and "PPT/PDF" Expectations
Well-packaged deliverables reduce ambiguity and prevent misuse.
Decision Tables (Genotype → Pass/Fail/Retest Rules)
Decision tables should encode genotype states, acceptance thresholds, and exceptions. Include clear rules for retesting borderline samples and for invoking phenotypic confirmation.
QC Summary Blocks That Prevent Misuse of Markers
A single-page QC summary should report per-sample and per-marker metrics, replicate concordance, batch comparability plots, and—when sequencing-based—imputation quality. This guards against uncritical reuse of markers outside their validated scope.
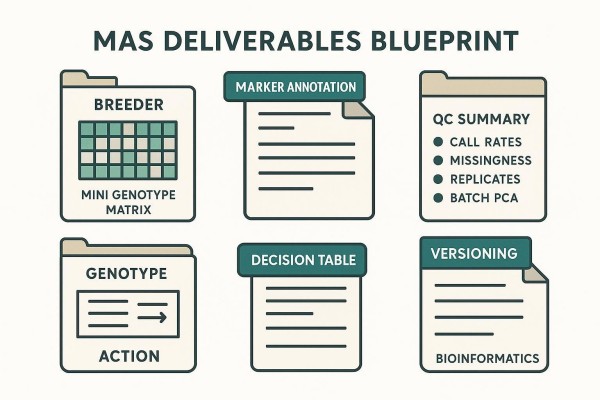
Reporting Template Elements That Make Results Reusable
Include a genotype matrix, marker annotation notes (effect/diagnostic allele, assay details, expected performance domain), QC summary, the decision table, and versioning notes indicating updates over time. Layouts designed for PPT/PDF allow program managers and breeders to reuse the materials in reviews and handoffs without losing essential context.
How CD Genomics Supports MAS Workflows
Service providers can support MAS projects across discovery, validation, and routine screening by delivering decision-ready genotypes, marker annotations, and QC summaries for research use only (RUO). For example, CD Genomics offers crop-focused SNP panels such as the Rice SNP Panel designed to aid marker-informed selection in rice breeding programs. Stage-based support typically spans assay design, high-throughput genotyping across array or sequencing-based platforms, and packaging outputs as genotype matrices, QC summaries, and genotype-to-action tables suitable for downstream decision-making.
FAQ
What is marker assisted selection, and how is it different from genomic selection?
- Marker-assisted selection targets one or a few known loci using diagnostic or tightly linked markers to identify individuals carrying favorable alleles, enabling earlier selection than phenotype alone when traits are slow or costly to measure. Genomic selection estimates breeding values using genome-wide marker data and is often preferred for highly polygenic traits where many small-effect loci govern performance.
What are the marker assisted selection steps from trait definition to a breeding decision?
- The practical sequence is: define the trait and decision unit, link evidence to markers through mapping or association, design assays and genotype with appropriate controls, apply QC gates and acceptance thresholds, and translate genotypes into decision rules that specify advance, retest, or redesign triggers.
How does genetic mapping and marker assisted selection connect—do I need mapping before I can use MAS?
- Mapping or association studies establish the evidence that a marker tracks the biology of interest. Programs can sometimes adopt previously validated markers from the literature, but internal validation in the target germplasm and environments is essential before routine deployment to ensure transferability.
When is marker assisted selection for disease resistance reliable enough to deploy across backgrounds?
- Disease-resistance markers should be diagnostic or extremely tightly linked, show consistent association in diverse germplasm, and demonstrate stable field performance across multiple environments or years. Programs should also verify concordance against legacy markers and document known limitations before widescale use.
How should marker assisted selection in animal breeding be designed to avoid confounding from relatedness or population structure?
- Animal programs should use models that account for genomic relationships, validate within families or using parent-wise cross-validation, and test transferability across breeds or lines when deployment will span multiple populations. These steps reduce the risk that apparent marker utility is an artifact of shared ancestry rather than biology.
Closing and Next Steps
The most reliable MAS programs elevate QC gates and decision rules to first-class citizens: markers are adopted only when the evidence ladder is climbed, thresholds are met, and deliverables make actions unambiguous. Teams can adapt the decision map and workflow here to their crop or species, calibrate thresholds to local constraints, and pilot MAS on traits where early genotyping avoids costly cycles. When external support is needed, stage-based services that provide decision-ready genotypes and QC summaries can streamline routine screening and reporting in a research-use-only framework.
References
- Paraschivu, M., et al. "Gene Pyramiding for Sustainable Crop Improvement against Biotic and Abiotic Stresses." Agronomy, vol. 10, no. 9, 2020, p. 1255. MDPI.
- Kumar, Sandeep, et al. "Recent Advancements in Molecular Marker-Assisted Selection and Its Applications in Plant Breeding Programmes." Plant Methods, 2021.
- Mardekian, S., et al. "Advances in Integrated Genomic Selection for Rapid Genetic Gain in Crop Improvement." Theoretical and Applied Genetics, 2022.
- Subedi, S., et al. "Genomic Selection: A Tool for Accelerating the Efficiency of Breeding in Plants and Animals." Frontiers in Genetics, 2022.
- Zhang, et al. "Genomic Selection in Plant Breeding: Key Factors Shaping Two Decades of Practical Application." The Crop Journal, 2024.
- Heaton, Matthew P. "A Primer on Sequencing and Genotype Imputation in Cattle." Journal of Animal Science, 2025.
- Stoll, Martin, et al. "Marker-Assisted Selection in Breeding for Fruit Trait Improvement: A Review." International Journal of Molecular Sciences, vol. 24, no. 10, 2023, 8984.
 Send a Message
Send a MessageFor any general inquiries, please fill out the form below.

