Bovine Genotyping Array Services (Cattle)
CD Genomics provides bovine SNP array genotyping built for breeding-focused research—supporting genetic evaluation, pedigree construction, heterosis assessment, and genomic selection with standardized genotype data plus clear QC reporting. RUO only (not for diagnostic use).
Service Highlights

Bovine SNP Array Genotyping Overview (50K Cattle Panel)
Our cattle SNP array genotyping service for breeding provides standardized bovine genotypes for genomic selection and breeding research workflows. The Bovine 50K-class SNP array supports genetic evaluation, pedigree construction, heterosis assessment, and trait-focused studies, with QC reporting and organized genotype deliverables plus optional light bioinformatics checks. For research use only (RUO), not for diagnostic use.
This service delivers bovine SNP array genotyping based on a ~50K SNP panel designed for breeding research applications. The design concept emphasizes broad genome-wide distribution and marker content aligned to functional genes and SNP sets associated with breeding-relevant traits and genetic backgrounds. For breeding programs and genomics teams, the practical value is consistency: standardized genotypes across cohorts so you can compare animals, lines, and populations over time using the same marker content and the same QC fields.
When discovery-first sequencing is needed before array tracking, explore our Bovine Genome Sequencing. Learn more about our overarching capabilities in SNP Microarray Services.
What to prepare before you start (fit check inputs)
Applications of Cattle SNP Arrays in Breeding & Agrigenomics
Below are common ways breeding and genomics teams apply bovine SNP array genotypes in cattle research projects.
Performance evaluation
Use SNP genotypes to support breeding-focused evaluation studies where genomic information complements phenotypes and pedigree records. Common applications include cohort comparisons, grouping/stratification for analysis, and genomic-enabled evaluation research.
Genetic diversity analysis
Use genotype matrices to characterize diversity within and between cohorts, explore population structure, and assess relatedness patterns in multi-breed or crossbred groups—useful for conservation studies and breeding program monitoring.
Functional gene research
Use standardized cohort-wide genotypes to study genotype patterns around genes or regions of interest and to support functional interpretations in breeding-relevant research designs.
Genomic selection breeding
Integrate SNP array genotypes into genomic selection research workflows, such as training population development, candidate evaluation, and genomic prediction studies. Optional light structure/relatedness checks can help confirm cohort readiness for downstream modeling (project-dependent).
Genetic defect and disease-resistance screening
Use SNP genotypes to support breeding research workflows involving known loci or marker sets associated with defects or resistance traits, when marker definitions are available and interpretation remains within a breeding research framework (not diagnostic use).
For related reading, see Advancing Bovine Parentage Assessment.
Key Features of the 50K Bovine SNP Chip (Coverage, Throughput, Customization)
Cost-efficiency for large-scale genotyping
The 50K-class array approach is commonly selected for large cohorts where consistent marker content and standardized outputs enable repeatable, program-level analysis across groups, seasons, or collaborating teams.
Customization flexible (supports SNP expansion)
If your program needs additional loci beyond the base array content (e.g., program-defined markers, emerging trait loci, population-specific variants), we can scope customization and SNP expansion when supporting inputs are available (such as candidate SNP lists and reference coordinates). Learn more about our Custom SNP Microarrays capability (project-dependent).
Convenient high-throughput capacity
We support projects designed for scaled genotyping with run-level capacity up to 2,304 samples per run, while maintaining consistent deliverables and QC fields across batches.
Accuracy (documented via QC fields and demo visuals)
Every project includes QC field reporting for practical usability assessment. In addition, demo visuals (below) show typical array documentation outputs such as chromosome-wide marker distribution, genotype cluster plots, and call-rate summaries for validation datasets of this array design.
End-to-End Workflow for Bovine SNP Array Genotyping
⚡ Bovine SNP array genotyping projects follow a clear, documented service flow:

Sample Intake → DNA QC → SNP Array Genotyping → Genotype Calling → QC Review → Deliverables → (Optional) Light Bioinformatics Checks
Bovine SNP Array Data QC Metrics
QC reporting helps you judge whether genotype data is usable for your intended breeding research workflow. Supported by our Microarray Services framework.
Sample-level call rate field(s)
Included to assess individual sample performance across the array.
Missingness overview
Identifies patterns of missing data across markers or groups.
Replicate/consistency checks
Assessed when technical replicates are incorporated into the project design.
Cohort-level summaries & documented exclusions
Cohort-level QC summaries are provided to flag outliers, along with documented exclusions and reason codes for policy-level clarity (project-dependent).
Deliverables & Data Formats for Cattle Genotyping
Deliverables are packaged so your team can integrate genotypes into breeding research workflows with minimal reformatting.
Genotype matrix
Sample IDs aligned to your provided metadata.
QC report
Sample-level and cohort-level QC fields.
Methods/parameter notes
Describes file structure, key fields, and documentation logic.
Methods notes are written for practical use: what each file contains, how sample IDs map to metadata, and what QC fields mean. If you have a preferred downstream toolchain, note it during scoping so the handoff can be aligned.
Data Demo
Demo examples:



Optional Bioinformatics Add-ons
Optional light bioinformatics checks can be added when you need quick cohort-level readiness confirmation for downstream analyses.
Case Study: Using 50K Bovine Genotypes for Genomic Selection / Parentage Workflows
Citation
Naserkheil M., Bahrami A., Lee D., Mehrban H. (2020). Integrating Single-Step GWAS and Bipartite Networks Reconstruction Provides Novel Insights into Yearling Weight and Carcass Traits in Hanwoo Beef Cattle. Animals 10(10):1836. View Paper
Background: Growth and carcass traits are economically important in cattle breeding programs. The study used SNP-array-enabled association analysis to identify genomic regions linked to these traits and to support breeding research decisions grounded in genomic evidence.
Methods: The authors analyzed a Hanwoo beef cattle cohort using a weighted single-step GWAS approach that integrates genotypes, phenotypes, and pedigree information in one step, evaluating genomic windows associated with multiple traits. They also used network-based reconstruction to explore trait–gene relationships for biological interpretation.
Results: The paper reports multiple trait-associated genomic regions and presents Manhattan plots showing the proportion of additive genetic variance explained by genomic windows across chromosomes for each trait studied.
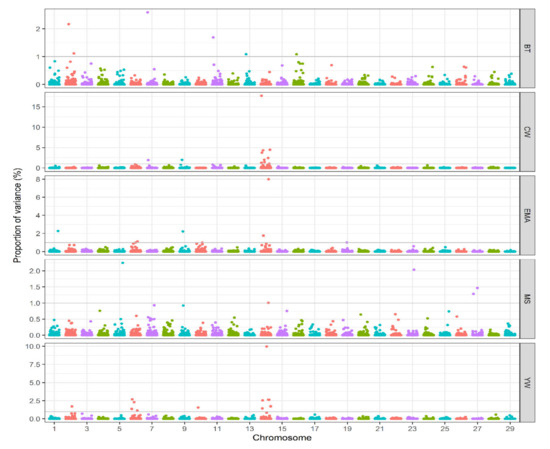
Manhattan plots of additive genetic variance explained by windows of adjacent SNPs across chromosomes for studied traits (from Naserkheil et al., 2020).
Conclusions: This published case illustrates how bovine SNP-array genotypes can be used to detect trait-associated genomic regions and support breeding research decisions when integrated with phenotypes and pedigree information using single-step association approaches.
Sample Requirements & Shipping for Bovine SNP Array Projects
| Item | Standard requirement (guideline) |
|---|---|
| Sample type | Genomic DNA (gDNA) |
| Species | Bovine (Cattle) |
| Volume | Sufficient volume for QC + genotyping (project-dependent) |
| Concentration | Provide consistent concentration across samples when possible |
| Purity | DNA suitable for array genotyping (avoid inhibitors) |
| Integrity | Intact gDNA recommended; avoid degradation |
| Labeling | Unique sample IDs matching your metadata sheet |
| Metadata | Sample ID, breed (if applicable), cohort/group, study goal, notes |
| Storage/handling | Maintain DNA integrity; avoid repeated freeze–thaw |
Submission checklist
FAQ: Bovine (Cattle) SNP Array Genotyping Service
Get a Quote
You can message us in any format—if helpful, use the checklist below to speed up routing and quoting.
References
Naserkheil M, Bahrami A, Lee D, Mehrban H. Integrating Single-Step GWAS and Bipartite Networks Reconstruction Provides Novel Insights into Yearling Weight and Carcass Traits in Hanwoo Beef Cattle. Animals. 2020;10(10):1836. https://doi.org/10.3390/ani10101836.
For research purposes only, not intended for clinical diagnosis, treatment, or individual health assessments.
 Send a Message
Send a MessageFor any general inquiries, please fill out the form below.
