Single-Cell Full-Length Transcriptome Sequencing
Single-cell full-length transcriptome sequencing (RNA-seq) is a technology that uses the 10x Genomics Chromium (10x) platform to obtain single-cell cDNA and combine it with transcriptome analysis on a long-sequence sequencing platform. This method combines the advantages of full-length transcriptome and single-cell sequencing, providing a powerful tool for discovering and quantifying isoforms, analyzing alternative splicing at the single-cell level.
Overview of Single-Cell RNA-Seq with Long-Read Sequencing
Single-cell RNA sequencing (scRNA-seq) has made up for the limitations of bulk RNA sequencing (RNA-seq). This method can resolve gene expression profiles in single cells for more accurate transcriptome assessment, and is currently receiving widespread attention and application. There are two main characteristics of the single-cell experiments, including the barcoding of cDNA synthesized by each cell with cell-specific barcodes and of individual mRNA molecules with molecular barcodes. Among them, the former method allows better quantification of transcripts at the single-cell level. Since molecular barcodes are usually degraded, it requires the high accuracy of the sequencing platform. ScRNA-seq is now a widely adopted approach for profiling transcriptomic heterogeneity. However, assessing transcript-level changes between cell types using current scRNA-seq protocols remains challenging, as they rely on short-read sequencing. Nowadays, long-read sequencing technology is widely used in scRNA-seq to generate complete transcript information in a single cell, overcoming the fundamental limitations of short-read sequencing.
Our Service Can Help Researchers Achieve
- Accurate identification of novel isoforms at the single-cell level.
- Isoform expression quantification at the single-cell level.
- High-quality alternative splicing analysis at the single-cell level.
Workflow of Our Services
10X Genomics captures 500-10,000 cells in a single sequencing run with extremely high cell throughput. Cells are labeled by 10X Genomics in a water-in-oil state, reverse transcribed into cDNA, then ligated by head-to-tail ligation process of the cDNAs and constructed into a long-read sequencing library of 10-15kb in length, which is then sequenced.

Analysis Pipeline and Contents
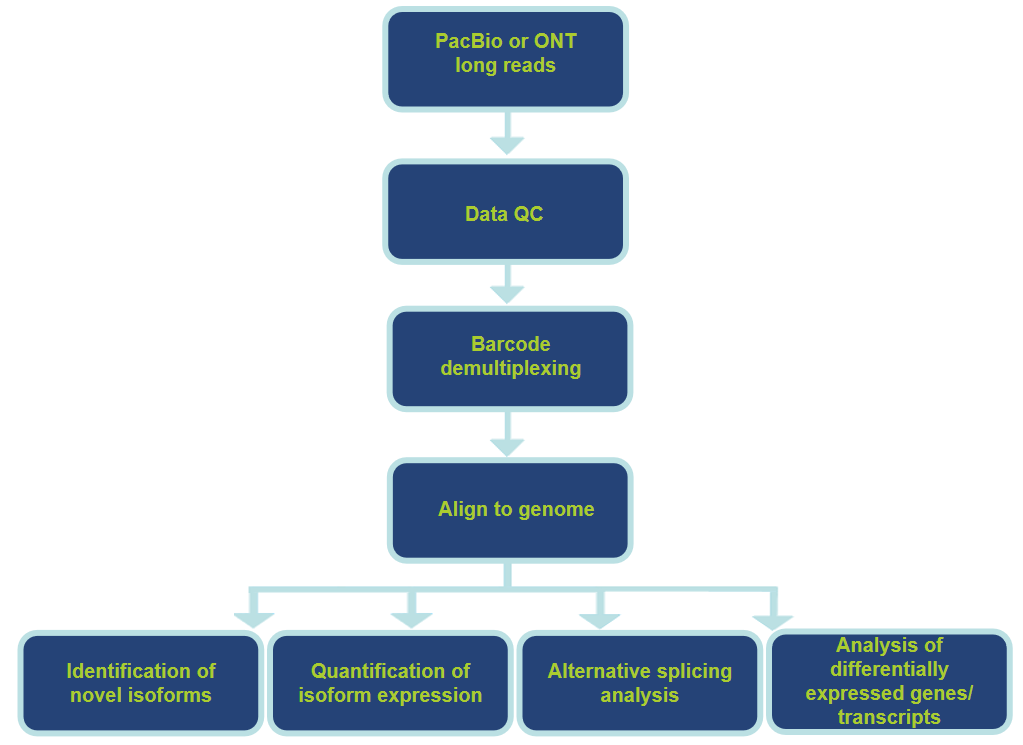
Features and Benefits of Our Services
- Eliminate the PCR Amplification Step: Say goodbye to PCR amplification during sequencing, which eliminates PCR bias and preference. View CD Genomics Platforms.
- Unprecedented Long Read Length: Our technology empowers you to sequence full-length transcripts, unlocking a deeper understanding of genetic information.
- A Paradigm Shift in Gene-Level Research: Move beyond traditional gene-level exploration and gain access to a wealth of data about full-length isoforms.
- Precision in Cell Barcode and Molecular Barcodes Detection: Our service ensures accurate detection of cell barcodes and molecular barcodes, providing high-quality data for your analysis.
- Comprehensive Resolution of Genetic Variations: We excel in resolving AS, mRNA point mutations, and fusion genes in single cells, giving you a comprehensive view of genetic complexity.
- No Dependency on Short Reads: You no longer need NGS data or short reads as a reference, simplifying your workflow.
- State-of-the-Art Bioinformatics Analysis: Our cutting-edge bioinformatics data analysis provides you with valuable insights and actionable information.
- High-quality library preparation and long-read RNA sequencing.
- Cutting-edge bioinformatics data analysis.
- Customized service to customer satisfaction.
- Customer service representatives are on call 24 hours a day, Monday through Friday.
At CD Genomics, we offer two strategies for our customers to choose from, including Pacific Biosciences (PacBio) and Oxford Nanopore Technologies (ONT) single-cell full-length transcriptome sequencing. Both methods have their own advantages in the field of RNA-Seq. Based on advanced platforms and years of experience in transcriptome analysis, we are committed to providing the best strategy to meet the project needs of our global clients. Our single-cell full-length transcriptome sequencing service has many applications. In addition to normal developmental and physiological studies, we focus on disease studies, such as tumor research and viral infections. If you have additional requirements or questions about our services, please feel free to contact us.
References
- Oikonomopoulos, S., et al. (2020). "Methodologies for transcript profiling using long-read technologies." Frontiers in genetics, 606.
- Tian, L., et al. (2021). "Comprehensive characterization of single-cell full-length isoforms in human and mouse with long-read sequencing." Genome biology, 22(1), 1-24.
- Wang, Q., et al. (2021). "Single-cell transcriptome sequencing on the Nanopore platform with ScNapBar." RNA, 27(7), 763-770.
For research purposes only, not intended for personal diagnosis, clinical testing, or health assessment