Nanopore Sequencing 101: Library Construction
At a glance:
- Importance of Effective Library Building
- Impact on Downstream Analysis
- Library Construction in Nanopore Sequencing
- DNA or RNA?
- Whether to PCR?
- Whether to add barcodes?
Library construction is indeed a critical step in the DNA sequencing process, especially in nanopore sequencing, as it directly impacts the quality and accuracy of the sequencing data obtained. The quality of the library will ultimately determine the success of the sequencing experiment and the reliability of the results.
Importance of Effective Library Construction
- Read Length and Accuracy: The success of nanopore sequencing lies in generating long reads with high accuracy. Effective library construction ensures that DNA or RNA fragments of the appropriate length are prepared for sequencing. Longer fragments lead to longer reads, which can be advantageous for various genomics applications, such as genome assembly and structural variant detection.
- Uniformity of Coverage: A well-constructed library results in even coverage across the genome or transcriptome. Uneven coverage could lead to regions being overrepresented or underrepresented in the sequencing data, affecting the ability to detect genetic variations accurately.
- Avoiding Bias: Biases introduced during library construction can impact the representation of certain DNA or RNA sequences in the final data. Library preparation protocols should aim to minimize biases, ensuring that all regions of interest have a fair chance of being sequenced.
- Adaptor Ligation Efficiency: Proper ligation of sequencing adapters is essential for the nanopore sequencer to identify the individual DNA or RNA molecules and obtain accurate sequence information. Ineffective adapter ligation can lead to read failures or shorter read lengths, reducing the overall quality of the sequencing data.
- Minimizing Contamination: Effective library building protocols help minimize the risk of contamination during the sequencing process. Contaminants from the laboratory environment or sample impurities can lead to misleading results or reduced data accuracy.
- Optimized Sample Handling: Proper library construction includes various sample handling steps, such as DNA or RNA extraction and fragmentation. Each of these steps must be optimized to ensure minimal sample degradation and preservation of the native genetic information.
- Quality Control: Thorough quality control during library construction allows researchers to identify any issues early on and take corrective measures. It helps prevent the propagation of errors through the sequencing process and ensures that only high-quality libraries are subjected to sequencing.
Impact on Downstream Analysis
The quality of the library directly affects the downstream analysis of sequencing data. A poorly constructed library can lead to several issues, including:
- Reduced sensitivity in detecting genetic variations and mutations.
- Ambiguity in identifying structural variations in the genome.
- Lower accuracy in assembling complex genomes or metagenomic samples.
- Difficulty in distinguishing true biological signals from technical artifacts.
In contrast, an effective library construction process ensures robust and reliable sequencing data, providing a solid foundation for meaningful biological insights and research discoveries.
Library Construction in Nanopore Sequencing
Nanopore library construction is indeed a relatively simpler and more streamlined process compared to traditional second-generation sequencing methods like Illumina sequencing. As you mentioned, the key advantage of nanopore sequencing is that it allows DNA or RNA molecules to pass through a nanopore, and the bases are identified by reading the electrical signals generated during this process. This eliminates the need for several laborious and error-prone steps involved in library construction for second-generation sequencing platforms.
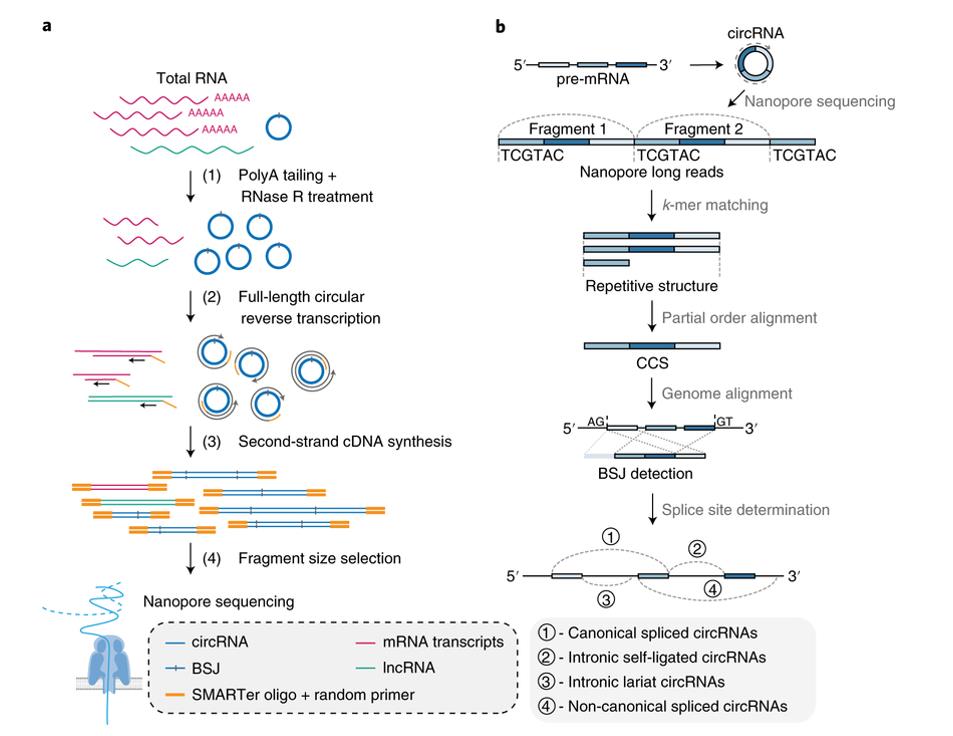 Workflow of bioinformatics analysis – CD Genomics
Workflow of bioinformatics analysis – CD Genomics
Nanopore Library Construction Steps:
- Fragmentation: The process begins with the fragmentation of DNA or RNA molecules into smaller segments. This can be achieved through mechanical shearing or enzymatic digestion, depending on the sample type and requirements. The goal is to obtain fragments of suitable length for nanopore sequencing.
- End Repair and A-tailing: The fragmented DNA or RNA ends may have uneven or damaged termini. To ensure uniformity and create sticky ends for further ligation, end repair enzymes are used to polish the ends. Following the end repair, adenosine (A) bases are added to the 3' ends, a process known as A-tailing.
- Adapter Ligation: Specific sequencing adapters are ligated to the A-tailed DNA or RNA fragments. These adapters contain essential signals for nanopore sequencing, such as motor protein recognition sites and electrical signal markers. The adapters facilitate the interaction of the DNA or RNA molecules with the nanopore and enable accurate sequencing.
- Formation of Y Junctions: After adapter ligation, Y junctions are formed to improve the efficiency of translocating the DNA or RNA molecules through the nanopore. Y junctions help align the DNA strands during nanopore sequencing and ensure better data quality.
- Addition of Motor Proteins: Motor proteins, such as phi29 DNA polymerase, are added to the Y junctions. These proteins aid in guiding the DNA or RNA molecules through the nanopore at a controlled speed, allowing the electrical signals to be accurately read during sequencing.
- Sequencing: Once the library is prepared with Y junctions and motor proteins, it is loaded onto the nanopore sequencer. As individual DNA or RNA molecules pass through the nanopore, the electrical signals generated are used to identify the bases and decode the sequence in real-time.
The choice of library construction method in nanopore sequencing depends on various factors, including the type of sample (DNA or RNA), the specific research objectives, and the desired attributes of the sequencing data (e.g., speed, read length, accuracy). Here are some considerations for each of the questions you raised:
DNA or RNA?
The first decision to make is whether the study requires sequencing DNA or RNA. Library construction methods differ for DNA and RNA samples, as the protocols must be tailored to the nucleic acid type.
Assessment of DNA Quality
To ensure the extracted DNA is of high quality and suitable for nanopore sequencing, several parameters should be evaluated. The Nanodrop instrument is commonly used to measure the concentration of DNA and assess its purity by measuring the absorbance at specific wavelengths. The following ratios are essential for determining DNA quality:
- A260/A280: This ratio indicates the purity of DNA and should ideally be around 1.8. A value close to 1.8 suggests minimal protein contamination.
- A260/A230: This ratio reflects the presence of contaminants such as polysaccharides and polyphenols. A value between 2.0 and 2.2 indicates good purity and minimal contamination from substances like phenol or salts.
By ensuring high A260/A280 and A260/A230 ratios, you can verify that the DNA sample is free from significant protein and chemical contaminants.
Assessment of Fragment Size
- For nanopore sequencing, obtaining high molecular weight (HMW) DNA is crucial to achieve long read lengths. Assessing the fragment size distribution of the DNA sample is important to determine whether it meets the requirements for nanopore sequencing. Several methods can be used to assess the fragment size:
- Agilent Bioanalyzer/Tapestation: These systems use capillary electrophoresis to analyze DNA fragments and provide a detailed profile of fragment sizes.
- Pulsed-Field Gel Electrophoresis (PFGE): PFGE is a powerful method to separate large DNA molecules based on size using an alternating electric field. It is particularly useful for analyzing HMW DNA.
- Fragment Analyzer: The Fragment Analyzer system uses an automated capillary electrophoresis platform to analyze DNA fragment sizes accurately.
By using these techniques, researchers can evaluate the distribution of DNA fragment sizes and confirm the presence of long fragments, which are essential for successful nanopore sequencing.
Whether to PCR?
Polymerase chain reaction (PCR) is often employed during library construction to amplify the DNA or RNA before sequencing. PCR amplification can increase the amount of material available for sequencing but may introduce biases and errors.
PCR-free library preparation can be used to avoid PCR-related biases, but it may require more starting material and potentially result in a lower yield.
The necessity of PCR in second-generation synthesis-sequencing protocols is significant due to various steps involving splicing, clustering, and other processes that require multiple PCR operations. However, this reliance on PCR also imposes limitations on read lengths, preventing long-read sequencing beyond 1000 base pairs, as observed in Sanger and Illumina sequencing.
Conversely, nanopore sequencing offers a compelling advantage by eliminating the need for PCR altogether. This capability enables the sequencing of ultra-long read lengths. Additionally, PCR introduces amplification bias, which introduces errors into the data.
Despite the benefits of avoiding PCR, certain research purposes may still require its use. For instance, amplification may be necessary when dealing with small amounts of raw DNA. Similarly, for relatively short transcripts that need to be reverse transcribed into cDNA, PCR becomes essential. Moreover, when sequencing 16S with limited data, mixing and library construction are required, which also involve PCR. Fortunately, nanopore sequencing supports these processes, including mixing and library construction. Notably, the nanopore technology currently allows mixed sample libraries with up to 96 samples, providing flexibility in different research contexts.
Whether to add barcodes?
Barcoding, also known as indexing, allows the multiplexing of multiple samples in a single sequencing run. Each sample is tagged with a unique barcode, making it possible to distinguish and analyze them individually after sequencing.
Barcoding is particularly useful in studies involving multiple samples, as it allows cost-effective pooling and sequencing of different samples together.
Reference
- Zhang, Jinyang, et al. "Comprehensive profiling of circular RNAs with nanopore sequencing and CIRI-long." Nature biotechnology 39.7 (2021): 836-845.
Related Services
Oxford Nanopore Sequencing
Oxford Nanopore Pre-Made Library Sequencing
For research purposes only, not intended for personal diagnosis, clinical testing, or health assessment