Introduction to Exosomes
Exosomes are small extracellular vesicles (EVs), ranging from 30 to 100 nm in diameter, carrying heterogeneous cargo, including miRNA, mRNA, lncRNA, lipids, and proteins. Exosomes widely present in very large quantities in all eukaryotic cells, mediating intercellular communication after being released into the extracellular space. In humans, exosomes nearly exist in all kinds of biological fluids, including plasma, urine, cerebrospinal fluid, saliva, ascitic fluid, and amniotic fluid, engaged in essential biological processes.
The non-coding RNAs in exosomes are thought to be major contributors to the molecular regulations occurring in the recipient cells. Furthermore, those endogenous RNAs (mRNA, snRNA, lncRNA, etc.) participate in immune modulation, cancer proliferation, and neuronal activity, leading to tremendous potential as a biomarker resource for diagnostics and therapeutic drug development. Accurate identification of exosomal RNA is essential. Although the qRT-PCR method has been widely used in exosome studies, it is sensitive, specific, and available for low sample volumes. PCR-based assays are species-specific for RNA and are limited to high-quality reference genes, clearly time-consuming and laborious. Therefore, deep sequencing is becoming the choice of technology to fast and efficiently obtain the global landscape of exosomal RNAs and their biological roles.
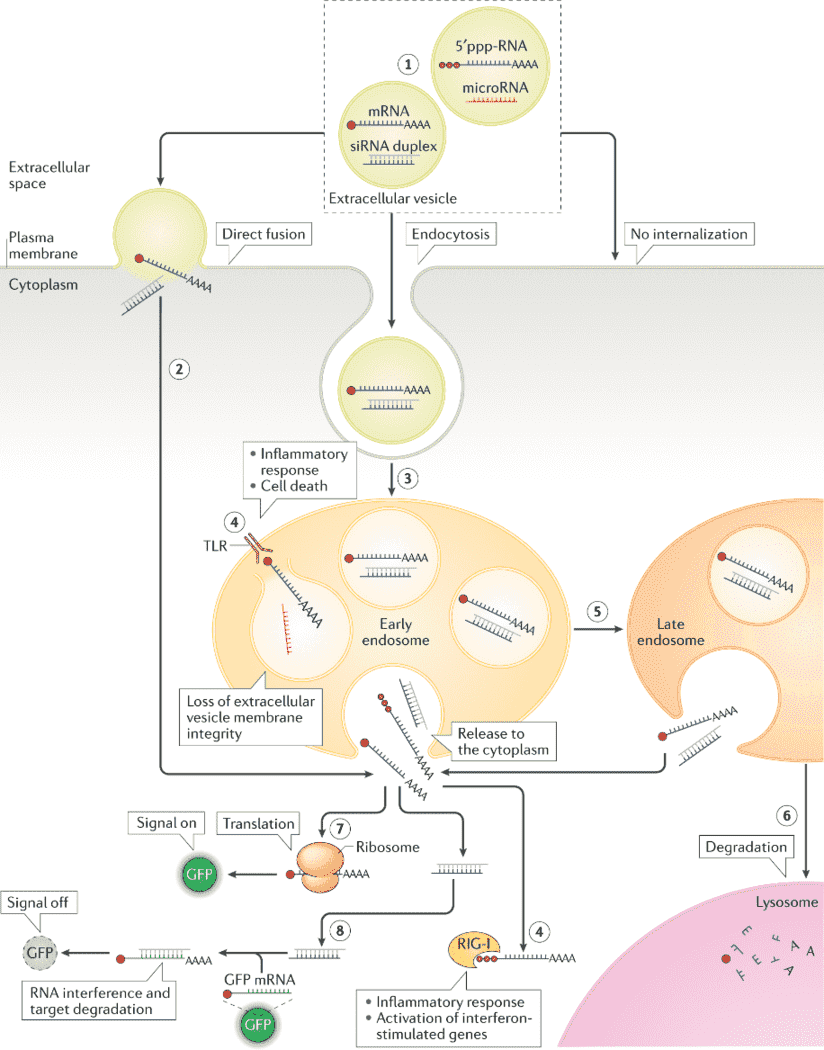
Figure 1. Extracellular vesicle RNA cargo interaction with recipient cells and its functional delivery (O’Brien K et al. 2020)
RNA Sequencing in Exosome Transcriptome Profiling
Cumulating data show that the RNA content of exosomes has interindividual variability and differs from that of their parent cells, which drives functional differences. However, quantification of the RNA cargo is quite challenged, due to the low numbers of cargo molecules per exosome (e.g., one miRNA per 100 exosomes, and one long RNA per 1,000). Massive RNA sequencing technologies have enabled reproducible characterization of nearly entire exosome RNA, either protein-coding or lncRNA transcripts, which is consistently applied to demonstrate the presence of complex RNA cargoes within exosomes populations released by various cells.
Low input RNA sequencing, for instance, advances scientists to investigate exosome-enriched or total RNA with minimal bias, even down to the nanogram level.
Full-length RNA sequencing has the advantage of producing full-length transcripts in a short time, without laborious assembly. It is quite suitable for identifying long transcripts (lncRNA, circular RNA, mRNA, etc.) and accurately annotating alternative splicing sites, fusion genes, and complex regions.
The Complete Workflow: from Isolation to Characterization of RNA
1. Exosome Isolation & Purification – Exosome extraction methods are mainly based on their physical characteristics and components, such as ultracentrifugation, density gradient flotation, ultrafiltration, chromatography, polymer-based precipitation, and immunoprecipitation. Some lipoprotein particles of similar size to exosomes in the sample may not be distinguished by those methods. Thus, there is still a challenge in extracting highly-pure exosomes. And the quality determines the suitability of characterizing various EV type-specific RNA cargo downstream.
2. Exosomal RNA Extraction & Purification – Another key part of ensuring the accuracy of exosome transcriptome analysis. Many approaches are developed for small and long RNAs, capture 5′-phosphorylated 3′-OH short RNAs, and whole transcriptome sequencing incorporates long RNAs more than small RNAs.
3. Library Construction
4. In-depth Data Analysis – Data process with exosome-seq mapping pipeline. Different sequencing solutions can profile exosomal RNAs about the expression levels and dynamics, understanding the physiological roles, with or without prior knowledge. As for long RNA, information about alternative splicing and gene fusion events can be revealed by long-read sequencing and complete annotation.
The Opportunities for Exosome RNA Sequencing
Thanks to high-throughput sequencing, we have novel insights into the picture of exosome RNAs in human or other species’ biofluids, where different extracellular RNA carriers can be exploited for biomarker discovery. With technical and biological limitations and challenges, considerations should be put into single-vesicle characterization, high concordant exosomal RNA quantification, etc. Further investigation of exosome transcriptome for monitoring dynamic progression may provide a novel view in developing RNA drug delivery systems and personal medicine.
References
- O’Brien K, Breyne K, Ughetto S, et al. RNA delivery by extracellular vesicles in mammalian cells and its applications. Nature reviews Molecular cell biology, 2020, 21(10): 585-606.
- Turchinovich A, Drapkina O, Tonevitsky A. Transcriptome of extracellular vesicles: state-of-the-art. Frontiers in immunology, 2019: 202.


 Sample Submission Guidelines
Sample Submission Guidelines