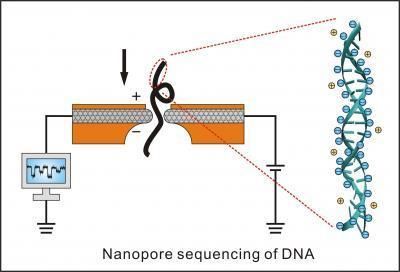
As a new technology of sequencing, the biggest advantage of nanopore is its low cost. Compared to other sequencing technology, such as sanger sequencing and next generation sequencing, the nanopore sequencing does not need any fluorescein for purification, and the process of DNA amplification is also unnecessary. It can save both money and time. Besides, the rapid reading speed is also one of the characteristics of nanopore sequencing. In the process of nanopore sequencing, DNA will go through the small channels which contain current. When the single nucleotide tries to get through the channel, it will try to stop the current and be read. However, the high-speed becomes a big problem here. It greatly reduces the precision and leads to some mistakes.
Recently, researchers from EPFL found a way to solve this problem. They identified a viscosity gradient system can be used to control the dynamics of DNA translocation through MoS2 nanopores, which greatly improved the precision of sequencing.
The researchers made a membrane which has a thickness of 0.7nm and 0.3nm-wide nanopores. Then the DNA was dissolved in the ionic liquids with high viscosity. Researchers can adjust the viscosity gradient system by changing the molecule of liquids. They slowed down the speed of DNA by this method.
If they can successfully test their research findings, it will lead to a revolution to the sequencing technology. Scientists have been searching for a way to reduce the cost of sequencing. It is well-known that sequencing plays a very important role in our medical research, and it has great potential in clinical application. However, the traditional method is costly and the time-consuming, which happens to be the advantage of nanopore sequencing. The new finding will not gain the cost or time of this technology, but it can make it more accurate and reduce the error. For some big sequencing projects like human genome sequencing, this will make a great change.


 Sample Submission Guidelines
Sample Submission Guidelines