In recent years, studies have been progressively conducted to assess the effect of distinct sampling techniques on the outcomes of fecal microbiome assessment, in order to address this unmet need in the area of microbiome research. These methodology researches are useful for researchers to create quite rational guidelines for the procedure of fecal sampling, handling, and sequencing.
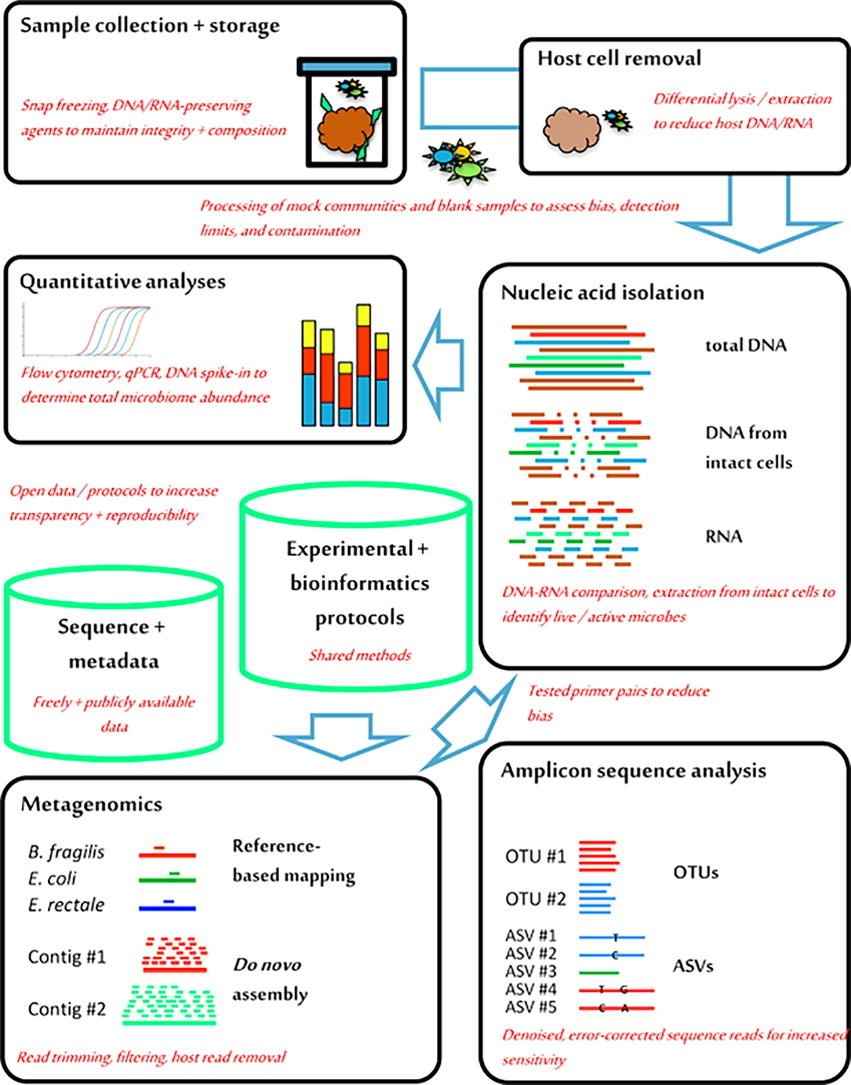
Figure 1. A brief workflow from stool collection to sequencing. (Fricker, 2019)
Process Prior to the Collection of Stool Samples
Since there are various host and environmental variables that can affect our gut microbiome, a microbiome research layout should consider the compilation of microbiome-associated clinical databases. These variables involve age, gender, birth records, ethnicity, geography, occupation, anthropometry, dietary habits, usage of probiotics/prebiotics, food supplements, bowel habit condition, antibiotic use background, and other affiliated histories of medication and surgery. Even so, as the host and environmental factors linked with the gut microbiome are constantly revised at a fast rate, these database collections are possibly the least definitive segment of the standardization of the guidelines. Lately, experts have discovered that with host diurnal cycles, the intestinal microbiome sequence can change, and even pet ownership, as well as human interaction frequency, can affect the microbiome. Thus, the recording of all probable microbiome-related medical evidence could be inefficient because both effectiveness and data gathering practicability are vitally valuable. For example, although it has been confirmed that a short-term nutritional change can significantly alter the gut microbiome, in most medical conditions, it may not be feasible to retrieve nutritional documents closely and repeatedly. Thus, the hypothesis-driven method is the efficient method for metadata collection.
Stool Sample Collection
A crucial element of specimen acquisition is that stool can be collected and used as a substitute for the lower intestinal (distal colon) microbiome for the research of microbiomes of the gastrointestinal tract, where most host-related microbes exist. Stool specimens can be obtained via swabs or bulk compilation in many instances, such as clinical settings, and then instantly frozen at 20 °C or below to avoid alterations in the bacterial diversity until DNA extraction can be finished. However, constant access to a consistent temperature at 20 °C or lower may not be possible for specimens collected in remote locations or needing delivery.
The general concepts for the compilation of fresh stool specimens are:
1. Prevent frozen-thaw cycles: it is well established that the frozen-thaw cycle is the main element in causing stool specimen DNA deterioration, which can result in bad quality of metagenomic DNA and downstream skew assessment.
2. Prevent repeated fluctuations in temperature: intense changes in temperature are massive stress in stool specimens for many bacteria, which can destroy microbes and exacerbate the deterioration of DNA.
3. Reduce the transport time: the shipping period should be reduced to avoid unwanted microbial development and decline. The cold storage could broaden the transportation time acceptably.
Sequencing Methods
Via the transmission of the gut microbiota, microbiome study is progressively being translated into medical applications. Over the last couple of years, in an extremely fast expansion, numerous studies investigating bidirectional mystery interactions between fecal microbiota and human health have been released. This is connected to the growth of budget next-generation sequencing (NGS) innovation so that massive volumes of DNA fingerprint data can be easily acquired in a single instrumental run to decode the diverse bacterial community in the fecal specimen. Nevertheless, it is crucial to remember whether metagenomic assessment experiments are obtained from a dependable stool specimen preservation procedure before performing an expensive high throughput sequencing. As the fecal gathering kit, transport status, storage status, and process of DNA extraction all have some effects on the quality of the specimen, it would be recommended to cautiously maximize the fecal sampling pipeline to obtain large metagenomic DNA for unbiased NGS and bioinformatic analysis.
References
- Fricker AM, Podlesny D, Fricke WF. What is new and relevant for sequencing-based microbiome research? A mini-review. Journal of advanced research. 2019, 19.
- Wu WK, Chen CC, Panyod S, et al. Optimization of fecal sample processing for microbiome study—the journey from bathroom to bench. Journal of the Formosan Medical Association. 2019, 118(2).
- Song SJ, Amir A, Metcalf JL, et al. Preservation methods differ in fecal microbiome stability, affecting suitability for field studies. MSystems. 2016, 1(3).
- Dominianni C, Wu J, Hayes RB, Ahn J. Comparison of methods for fecal microbiome biospecimen collection. BMC microbiology. 2014, 14(1).


 Sample Submission Guidelines
Sample Submission Guidelines