Mitochondrial diseases (MitD) are an important clinical or medical challenge. Research has shown that genome sequencing can help to enhance the understanding of the genetic causes of MitD and to accelerate the development of functional research, clinical tests and therapeutic tools.
What is Mitochondrial Disease?
Mitochondria are membrane-bound organelles found in most eukaryotic cells that produce a majority of the chemical energy needed to power cellular biochemical reactions. The types of pathogenic variants of mitochondrial disease include pathogenic single nucleotide variants (pSNV), mitochondrial DNA (mtDNA) rearrangements and deletions, and altered mtDNA copy number (CN).
Although relatively rare, it is estimated that one in 3,500-6,000 people are affected by or at risk of developing mitochondrial disorders. High energy-consuming tissues are most susceptible, such as the brain, heart, muscles and eyes, but cases may present with a wide range of symptoms. Clinically relevant phenotypes include type 2 diabetes, multiple sclerosis, intellectual disability, and gastrointestinal disorders, etc. Mitochondrial disease can therefore be difficult to diagnose and distinguish from other diseases.
The Complexity of Human Genome
Human mtDNA, a 16,569 bp circular DNA molecule with no introns, is completely separate from the nuclear genome and is inherited maternally. It is estimated there are 1,500 proteins in the mitochondrial proteome, most of which are controlled by the nuclear genome. Mitochondrial disorders may therefore be caused by deleterious variants in mtDNA or nuclear DNA (nDNA). More than 300 nuclear genes are known to cause MitD, involved in assembly factors, mitochondrial structure, coenzyme Q biosynthesis, protein synthesis and mtDNA maintenance. These mutations can be inherited in any mode of inheritance: sporadic, maternal, autosomal dominant, autosomal recessive or X-linked.
In addition, the mitochondria themselves generate additional complexity. Studies have shown that the mtDNA mutation rate is much higher than that of nDNA, probably 10 times. This can occur throughout life at the germline level or at the somatic level, meaning that even if a specific variant is not detected in a tissue, it cannot be ruled out that the variant is not present in other tissues or organs.
Because of its diversity and complexity, MitD is difficult to diagnose and treat in clinic.
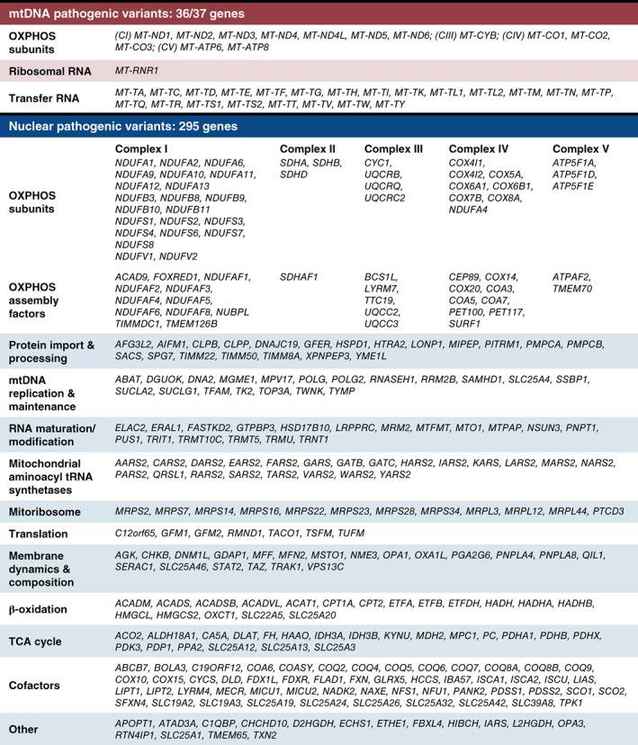
Figure 1. List of genes currently associated with mitochondrial disease (Thompson K et al., 2020)
Whole Genome Sequencing in Mitochondrial Disease Research
The genetic heterogeneity and potential involvement of both genomes means that MD is not well suited to targeted genome sequencing approaches. Whole genome sequencing has the advantage of covering almost all of the nuclear and mitochondrial genomes in a single test and obtaining information on single nucleotide variants (SNV), short insertions and deletions and structural variants (SV). It also allows for excellent sequencing depth. Whole genome seqeuncing is known to detect levels of heterogeneity down to 1% and can identify variants in blood samples of most patients, reducing the need for subsequent tissue biopsy. With the costs of sequencing is decreasing, genome sequencing is likely to become the method of choice for MitD genetic mutation identification, functional research and molecular diagnostics.
References
- Rius R, Compton AG, Baker NL, et al. Application of Genome Sequencing from Blood to Diagnose Mitochondrial Diseases. Genes, 2021, 12(4): 607.
- Thompson K, Collier JJ, Glasgow RIC, et al. Recent advances in understanding the molecular genetic basis of mitochondrial disease. Journal of inherited metabolic disease, 2020, 43(1): 36-50.
- Riley LG, Cowley MJ, Gayevskiy V, et al. The diagnostic utility of genome sequencing in a pediatric cohort with suspected mitochondrial disease. Genetics in Medicine, 2020, 22(7): 1254-1261.
- Schon KR, Ratnaike T, Van Den Ameele J, et al. Mitochondrial diseases: a diagnostic revolution. Trends in Genetics, 2020, 36(9): 702-717.
- Molnar MJ, Kovacs GG. Mitochondrial diseases. Handbook of clinical neurology, 2018, 145: 147-155.


 Sample Submission Guidelines
Sample Submission Guidelines