CD Genomics is now providing 2b-RAD sequencing which is a novel reduced-representation whole genome sequencing and restriction-site associated DNA (RAD) sequencing method for linkage mapping and determination of genome-wide variants in a cost-effective way.
Introduction to 2b-RAD
2b-RAD method uses type IIB restriction enzymes (REs), which share the feature of cutting the genomic DNA at both sides of the recognition to produce a fixed-size dsDNA fragment and result in protruding, noncohesive ends. Subsequently, DNA fragments of interest are captured by a biotinylated adaptor specific to the initial enzyme. Compared with 2b-RAD, similar methods like restriction-site associated DNA (RAD) and genotyping-by-sequencing (GBS) technologies lack the specificity to regions of interest, and produce many sequences originated from non-informative and repetitive regions.
Sequencing of these short and uniform tags brings many benefits, including targeting of all restriction sites, even sequencing depth across sites, high reproducibility for quantitative measurement, low sequencing cost per tag and low sensitivity to DNA degradation. To benefit from the gradually increased sequencing capacity of next-generation sequencing (NGS) platforms, we also combine an advanced protocol that allows the preparation of five concatenated isoRAD tags for Illumina paired-end (PE) 150 bp sequencing, which provides researchers more power and flexibility in devising effective library configurations to meet specific research purposes.
Applications
- Bin Map construction and QTL location
- Population genetic study
- Population evolution analysis
- Genome-wide association study
- Genome phylogeny
- Genomic selection
- Linkage and association mapping
- Discrimination of microbial strains
- Detection of somatic mutations
Advantages of 2b-RAD
- Accurate and affordable
- Flexible tag number
- Consistent label length
- High density of markers
- Not requiring interim purification steps
- Allowing low DNA input and degraded DNA
- Highly reduced 2b-RAD libraries require much less sequencing for accurate genotyping
2b-RAD Workflow
The general workflow for 2b-RAD includes sample
preparation, 2b-RAD library preparation, high-throughput sequencing and bioinformatics analysis. Our highly
experienced expert team executes quality management, following every procedure to ensure confident and unbiased
results. The library construction for 2b-RAD consists of four major stages (BsaXI digestion, ligation,
amplification and barcoding).

Service Specifications
Sample Requirements
|
|
Sequencing
|
|
Bioinformatics Analysis
|
CD Genomics is proud to offer advanced 2b-RAD sequencing technology. Our service covers the complete process, from quality control of genomic DNA to determination of genotypes and population diversity. If you have additional requirements or questions, please feel free to contact us. CD Genomics is the licensed service provider for this technology. Keygene N.V. owns patents and patent applications protecting its Sequence Based Genotyping technologies.
1. What are the advantages of 2b-RAD?
CD Genomics is proud to offer advanced 2b-RAD sequencing technology, which harbors numerous advantages over original RADseq and GBS. The comparison between different reduced representation whole genome sequencing methods is outlined in Table 1.
Table 1. Technical comparison between 2b-RAD and other NGS-based reduced representation genotyping methods.
Technology |
RAD-seq, ddRAD | GBS | 2b-RAD |
| Library Construction | Complex, including sonication, fragment selection and multiple steps of DNA purification | No sonication and fragment selection steps, end-repair | Streamlined, no sonication and fragment selection steps, end-repair |
| Fragment Size | Non-uniform | Non-uniform | Uniform |
| Tag Density Adjustment | Difficult | Possible but not tested | Flexible |
| Tag Sequencing Depth | Highly divergent depth between different tags | Highly divergent depth between different tags | Uniform sequencing depth between tags |
| Data Analysis | False positive caused by repetitive sequences | False positive caused by repetitive sequences | Remove the distraction of repetitive sequence by using iML |
2. What are the disadvantages of 2b-RAD?
Although 2b-RAD is a powerful method for high-throughput genotyping, it also has some drawbacks. 2b-RAD works only on diploid species. Additionally, the short tags may not be long enough for locus discrimination in complex genomes.3. How is the 2b-RAD library prepared?
The library construction for 2b-RAD consists of four major stages (BsaXI digestion, ligation, amplification and barcoding). Following DNA digestion by BsaXI, adaptors are ligated to the fragments through cohesive-end, and specific barcodes are incorporated into each sample through PCR amplification using degenerated linkers. Then the single-tag constructs are produced using modified adaptors and biotin-labeled primers, which can be further digested by SapI to generate distinct cohesive ends and then ligated in a predefined order to produce five concatenated tags. Samples are then pooled and sequenced using Illumina technology.
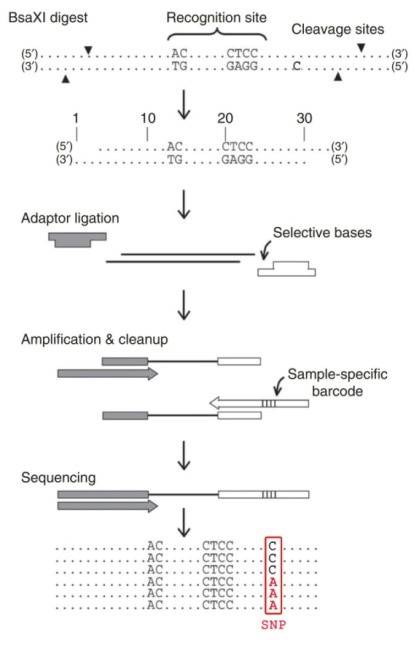 Figure 1. Schematic overview of the
2b-RAD procedure.
Figure 1. Schematic overview of the
2b-RAD procedure.
4. Can 2b-RAD detect SNP in repetitive regions?
2b-RAD may not detect SNP in repetitive regions that include a small number of restriction sites.
5. How many individuals are required for the construction of linkage map?
At least more than 100 individuals are necessary for constructing genetic linkage map.
6. For natural populations, how many samples are required?
Natural populations often have more SNP. The number of samples depends on the phenotype of interest, ranging from tens of samples to hundreds. The more samples you submit, the more accurate genotyping results you get.7. What is the requirement for species?
Diploids with or without a reference genome are suitable for 2b-RAD. However, polyploids for 2b-RAD had better have a reference genome, and risks are still existing.
A high-resolution genetic linkage map and QTL fine mapping for growth-related traits and sex in the Yangtze River common carp (Cyprinus carpio haematopterus)
Journal: BMC Genomics
Published: 2018 April 2
Abstract
The authors constructed a high-resolution genetic linkage map by utilizing 7820 2b-RAD and 295 microsatellite markers in a F2 family of the Yangtze River common carp (C. c. haematopterus). They mapped a set of suggestive and significant QTLs for growth-related traits and sex on this linkage map. The genetic map and these QTL-derived candidate genes and markers are useful for further studies on the Yangtze River common carp.
Results
1. Construction of the high-resolution linkage map
A total of 8115 markers (7820 2b-RAD markers and 295 SSRs) were grouped into 50 LGs that was consistent to the haploid chromosome number by using the JoinMap 4.1 software.
 Figure 1. The sex-averaged genetic linkage map of the Yangtze River common carp C.c. haematopterus
constructed based on 2b-RAD and microsatellite markers.
Figure 1. The sex-averaged genetic linkage map of the Yangtze River common carp C.c. haematopterus
constructed based on 2b-RAD and microsatellite markers.
2. Comparative genome mapping
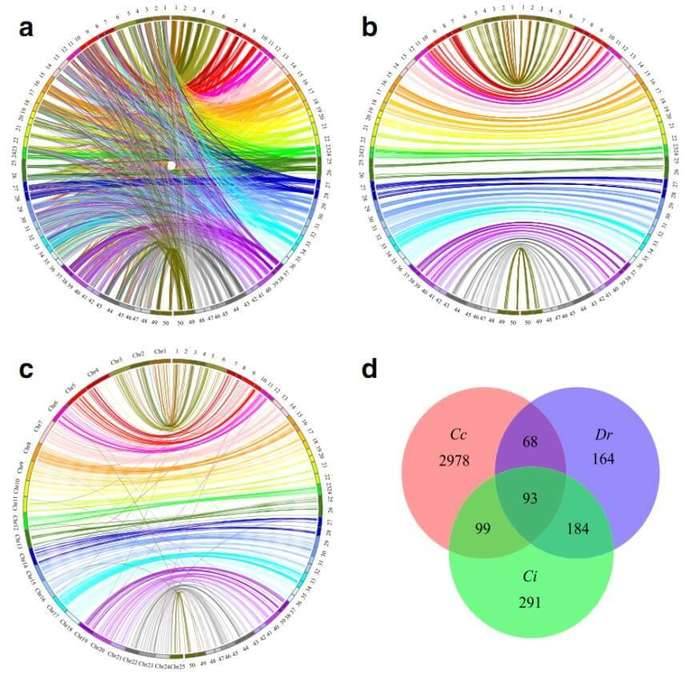 Figure 2. Circos diagram representing syntenic relationships between C. c. haematopterus (right) and (a
and b) C. c. carpio (left) and Danio rerio and (d) Venn diagrams describing overlaps among
uniquely aligned markers that mapped to genome of C. c. carpio (Cc), D. rerio (Dr) and C.
idellus (Ci).
Figure 2. Circos diagram representing syntenic relationships between C. c. haematopterus (right) and (a
and b) C. c. carpio (left) and Danio rerio and (d) Venn diagrams describing overlaps among
uniquely aligned markers that mapped to genome of C. c. carpio (Cc), D. rerio (Dr) and C.
idellus (Ci).
3. Fine QTL mapping for growth-related traits and sex.
A total of 21 QTLs associated with growth-related traits were detected on 12 LGs, including two genome-wide significant QTLs and 19 chromosome-wide significant QTLs.
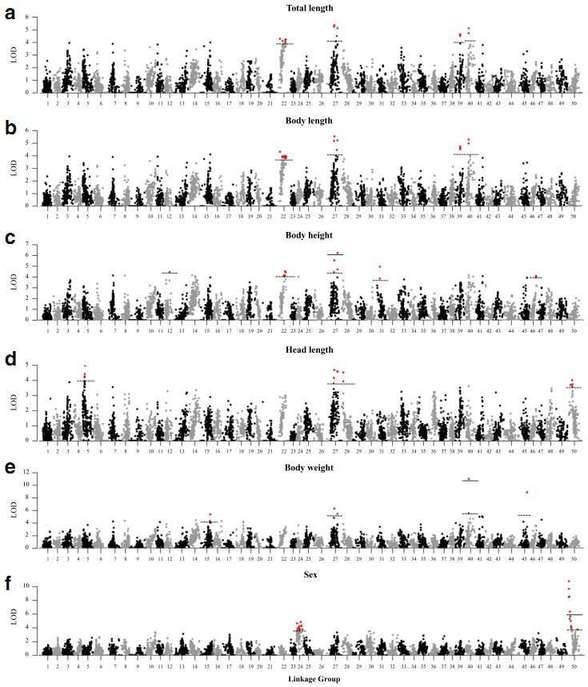 Figure 3. A genome scan of LOD profiles for (a) total length, (b) body length, (c) body height, (d) head length,
(e) body weight and (f) sex in C. c. haematopterus. The dashed and solid lines indicated the
chromosome-wide and genome-wide significance thresholds.
Figure 3. A genome scan of LOD profiles for (a) total length, (b) body length, (c) body height, (d) head length,
(e) body weight and (f) sex in C. c. haematopterus. The dashed and solid lines indicated the
chromosome-wide and genome-wide significance thresholds.
4. Potential candidate genes for growth
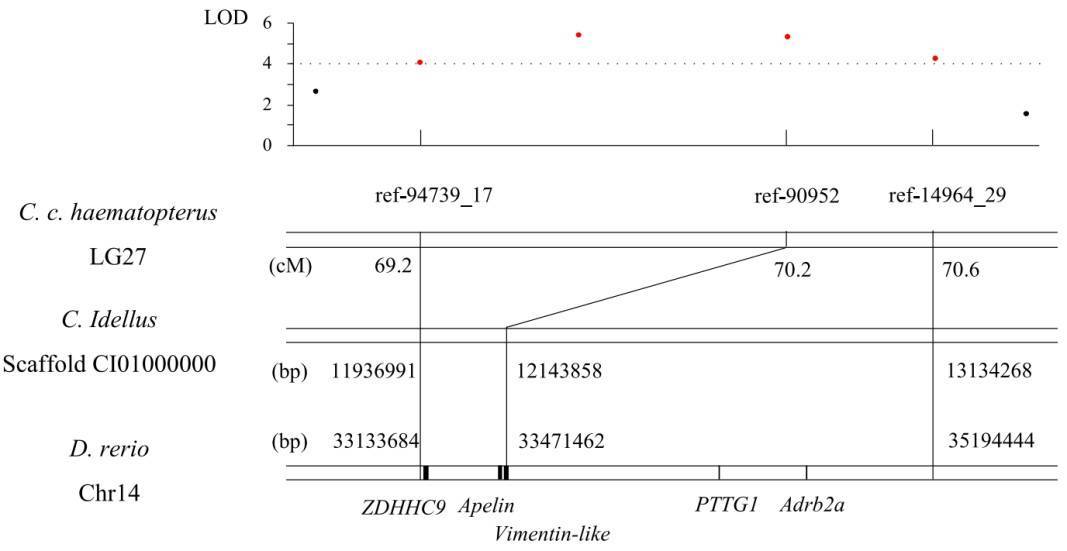 Figure 4. The QTL region for growth traits on LG27 of C. c. hasematopterus and its homologous region in
genomes of D. rerio and C. idellus.
Figure 4. The QTL region for growth traits on LG27 of C. c. hasematopterus and its homologous region in
genomes of D. rerio and C. idellus.
Reference:
Feng X, Yu X, Fu B, et al. A high-resolution genetic
linkage map and QTL fine mapping for growth-related traits and sex in the Yangtze River common carp (Cyprinus
carpio haematopterus). BMC genomics, 2018, 19(1): 230.


 Sample Submission Guidelines
Sample Submission Guidelines